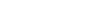Complex heatmap visualization
- 01 August 2022
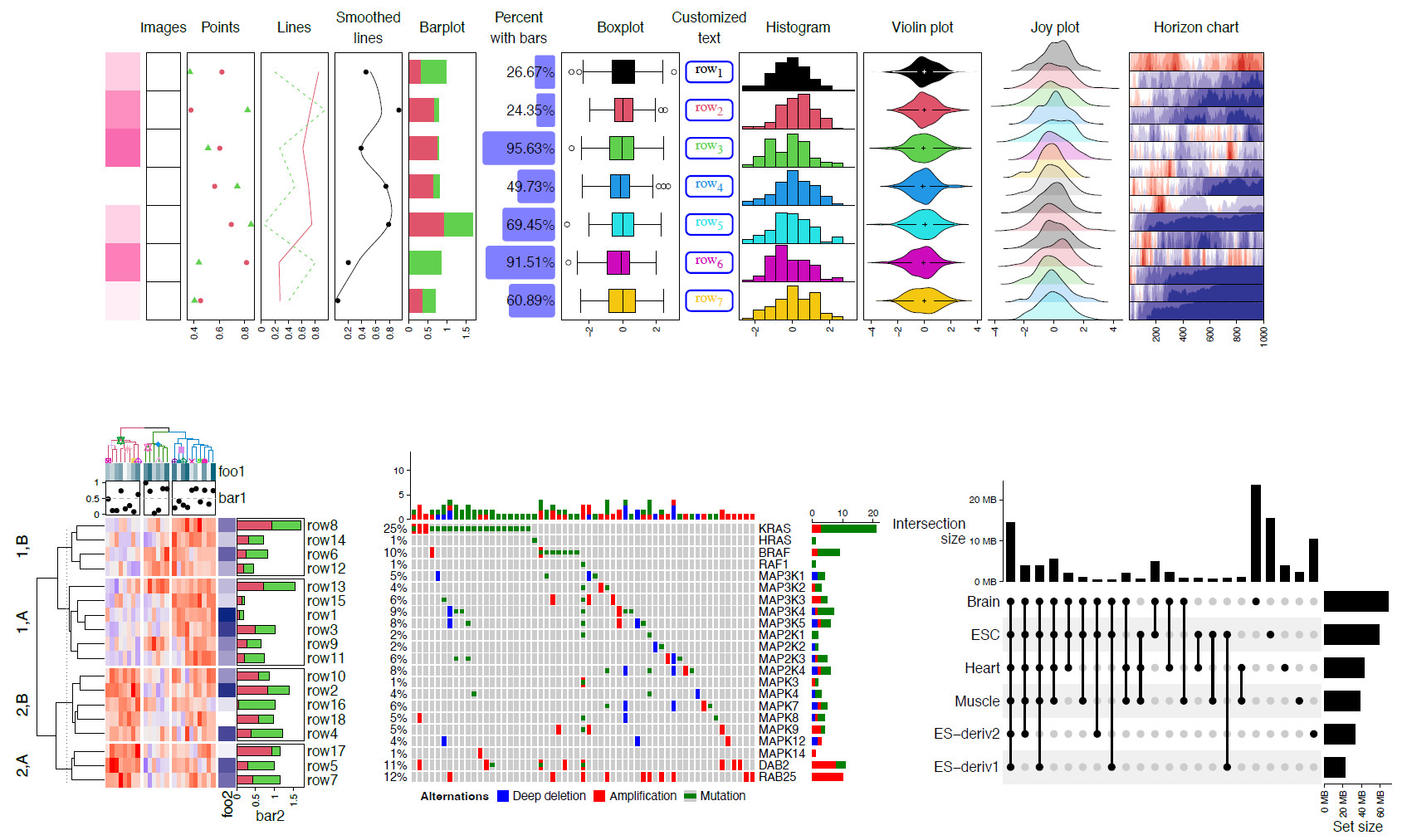
Complex heatmap is a powerful visualization method for revealing associations between multiple sources of information. We have developed an R package named ComplexHeatmap that provides comprehensive functionalities for heatmap visualization. It has been widely used in the bioinformatics community. We give a comprehensive introduction to the current state of ComplexHeatmap in this article.
A painless way to customize Circos plot: From data preparation to visualization using TBtools
- 04 July 2022
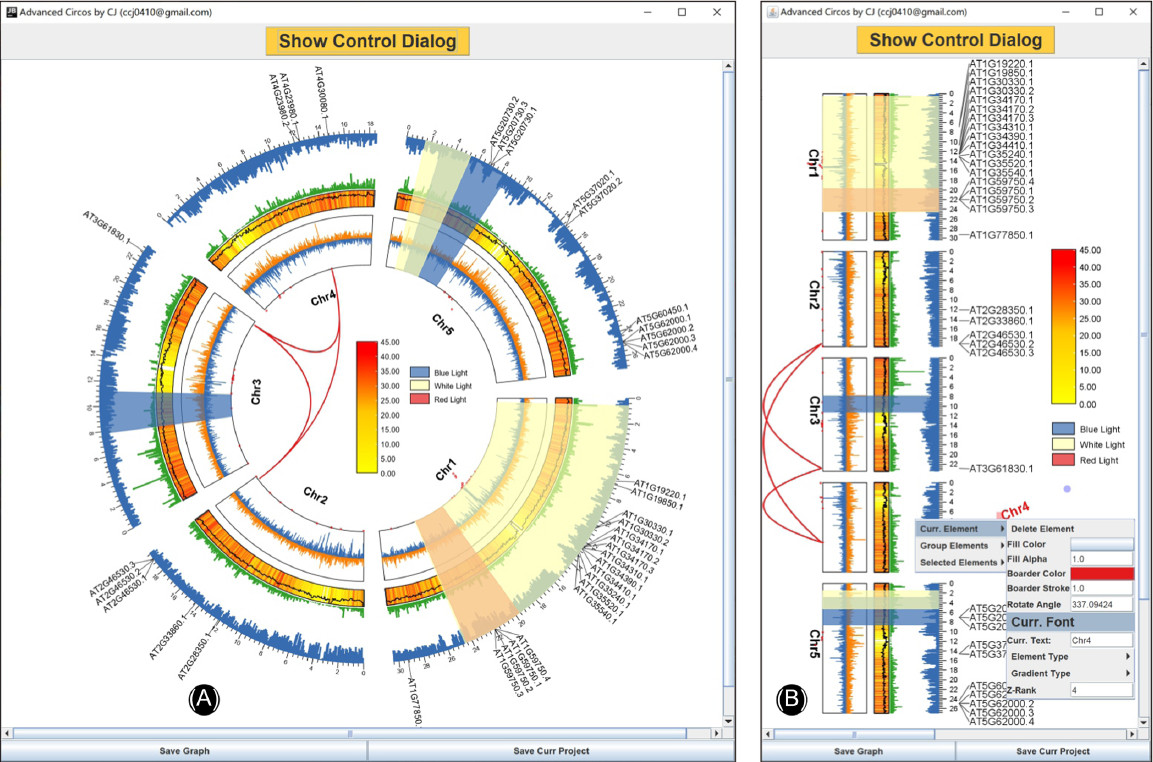
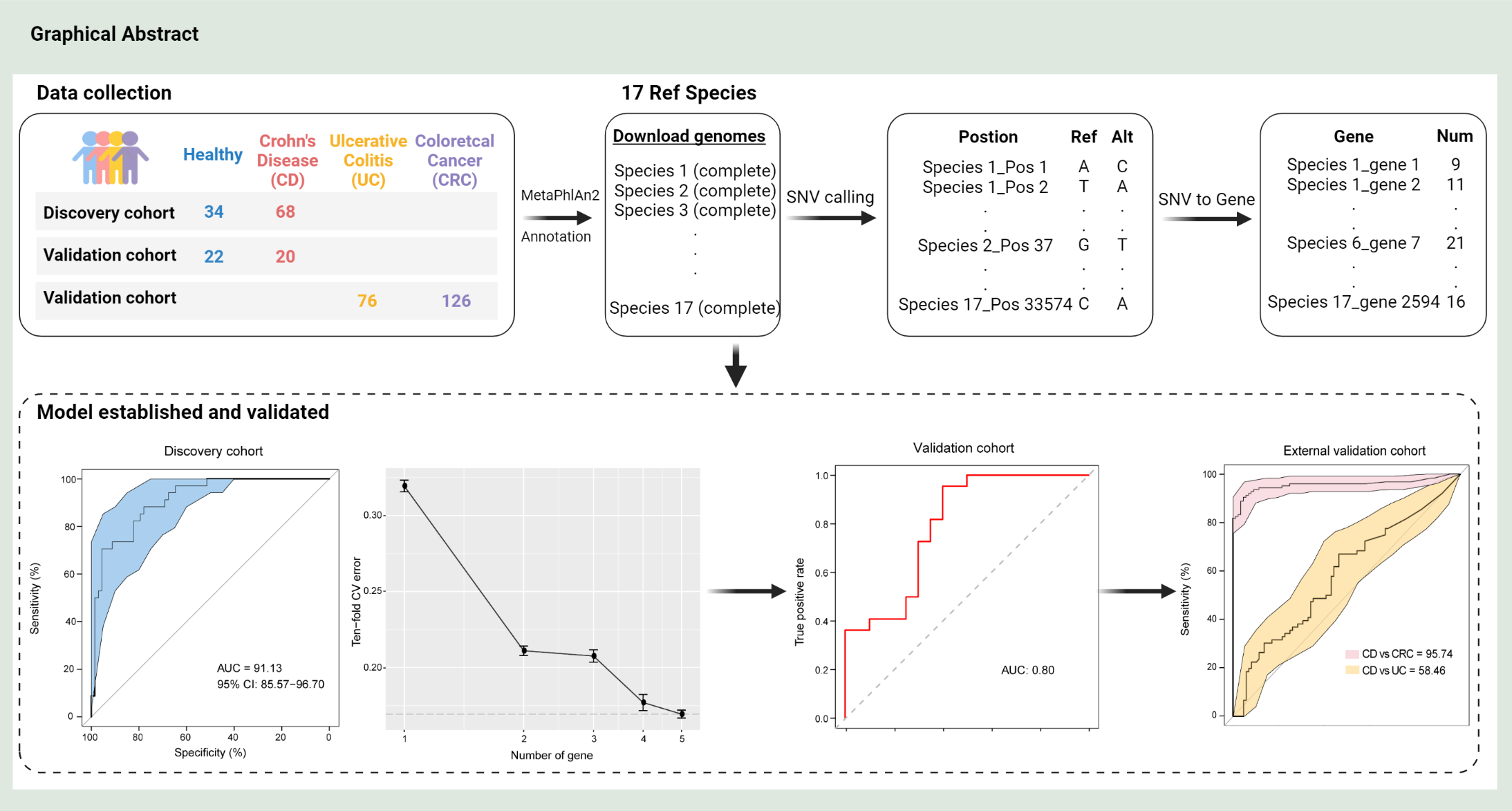
Establishing a novel inflammatory bowel disease prediction model based on gene markers identified from single nucleotide variants of the intestinal microbiota
- 24 July 2022
The human lung microbiome—A hidden link between microbes and human health and diseases
- 16 June 2022
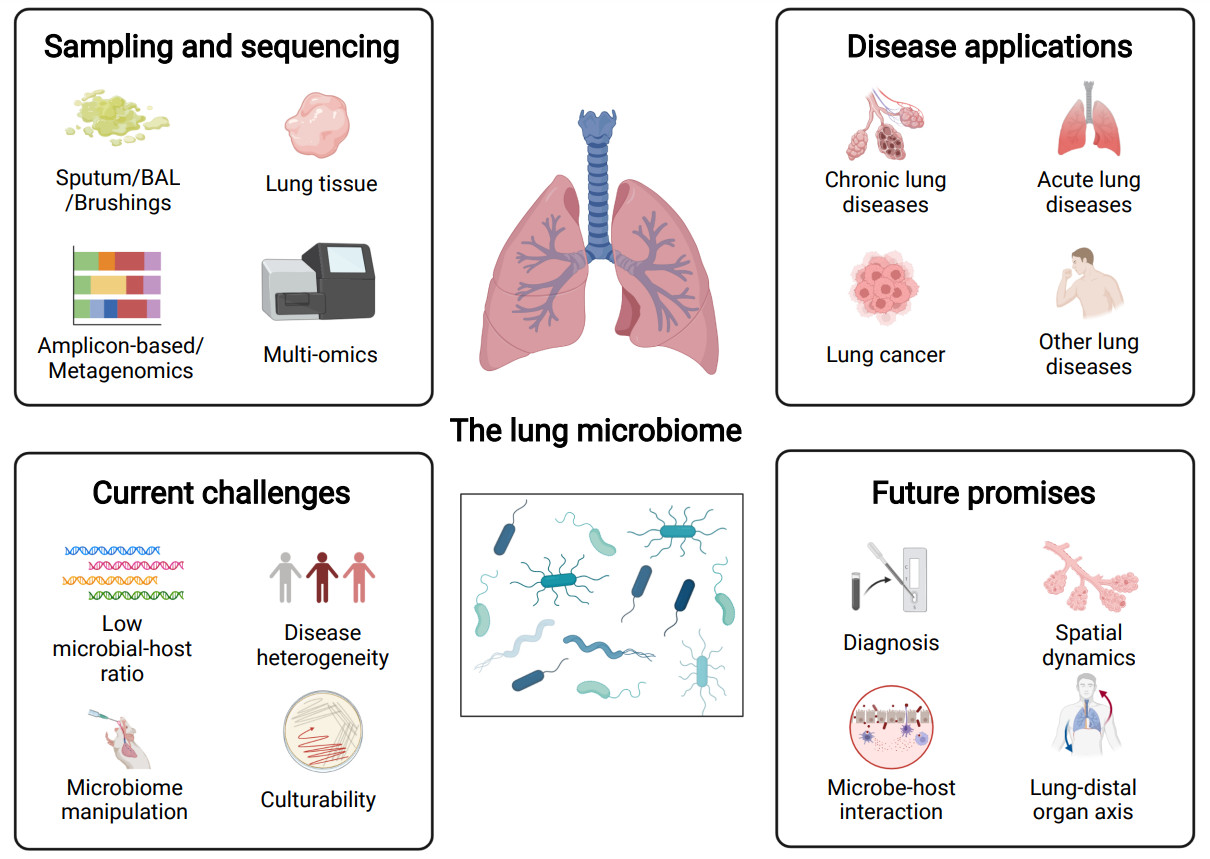
Sputum, bronchoalveolar lavage, bronchial brushing, and lung tissue are the routine sample types for the lung microbiome. Multiomics have been increasingly applied for characterizing the lung microbiome. Lung microbiome is broadly implicated in chronic and acute lung diseases, lung cancer, and other lung diseases, with potential mechanistic implications. Current challenges of the lung microbiome studies include low microbial-to-host ratio, high disease heterogeneity, and the difficulty in precisely manipulating and culturing the lung microbiome. Future potential topics for lung microbiome studies include understanding the diagnostic potential of the lung microbiome, its spatial dynamics, its mechanistic interaction with the host, and the crosstalk between the lung and distal organs.
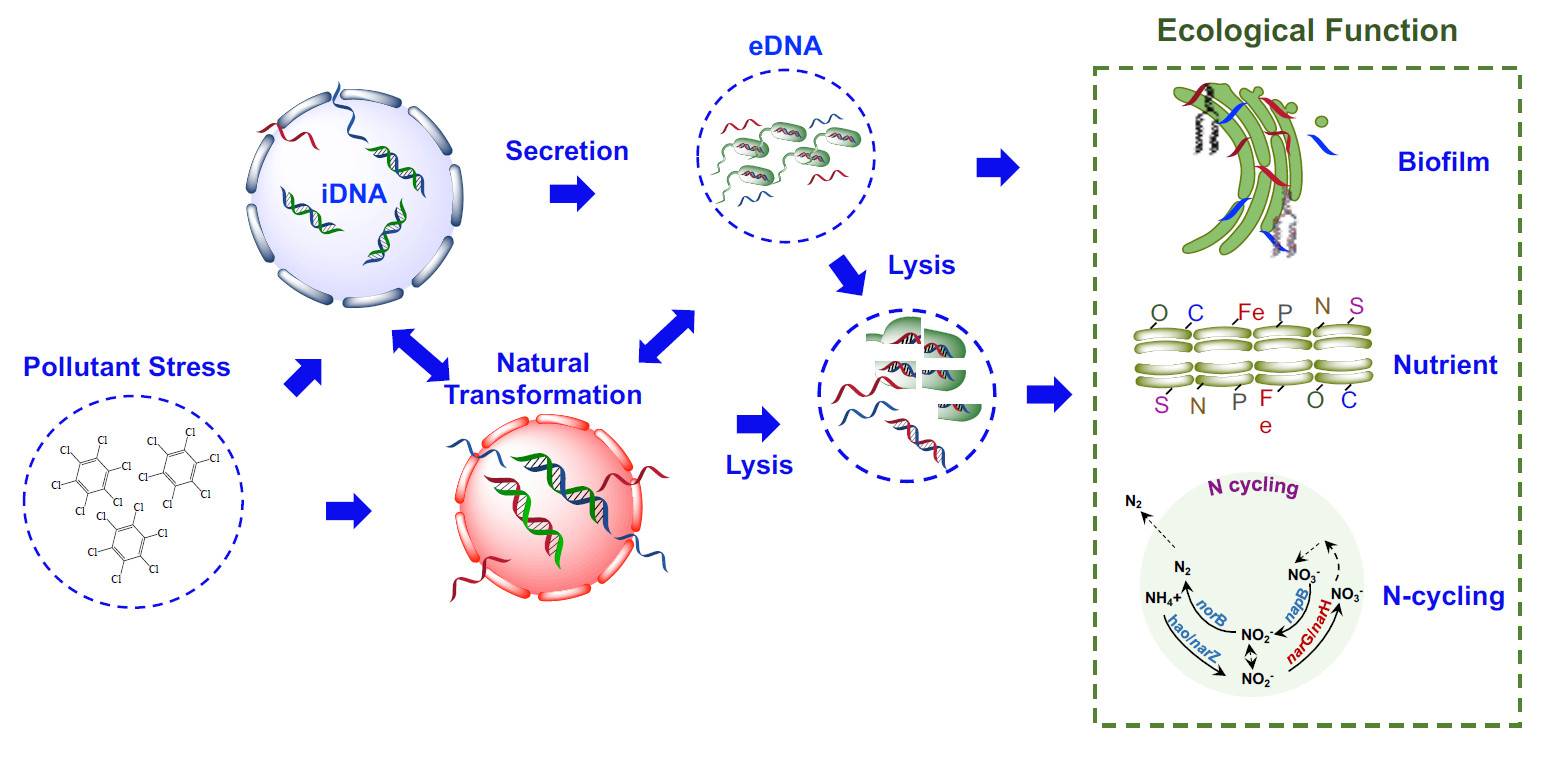
Dynamics, gene transfer, and ecological function of intracellular and extracellular DNA in environmental microbiome
- 20 June 2022
Are short-read amplicons suitable for the prediction of microbiome functional potential? A critical perspective
- 04 July 2022
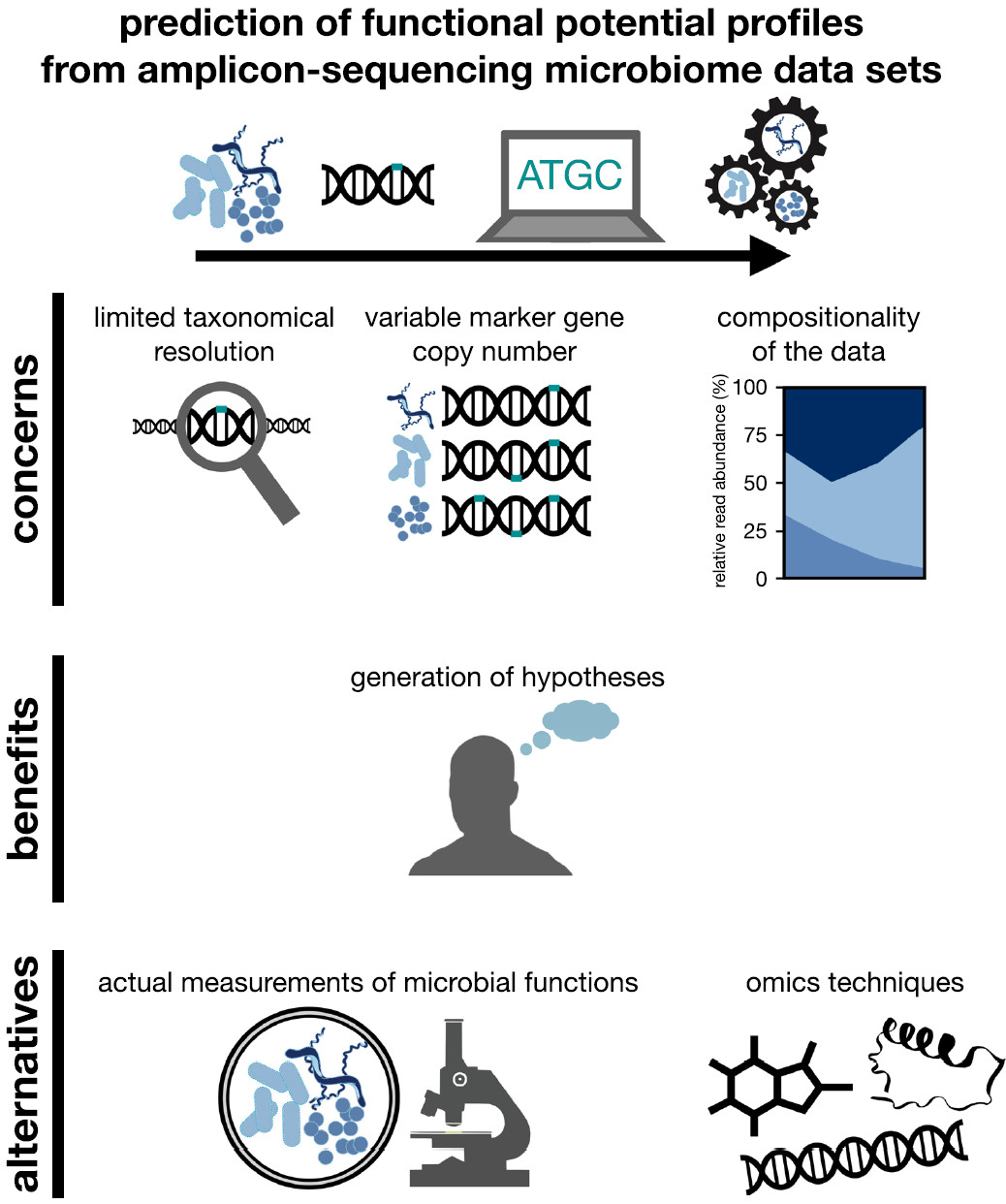
We raise concerns regarding the biological validity of predicting functional potential from microbiome taxonomic profiles generated by short-read amplicon sequencing. We reason that predicted functional potential profiles can be improved by employing long-read sequencing technologies, which in combination with independent measurements of actual functions can constitute a powerful approach to understanding microbiome functioning.
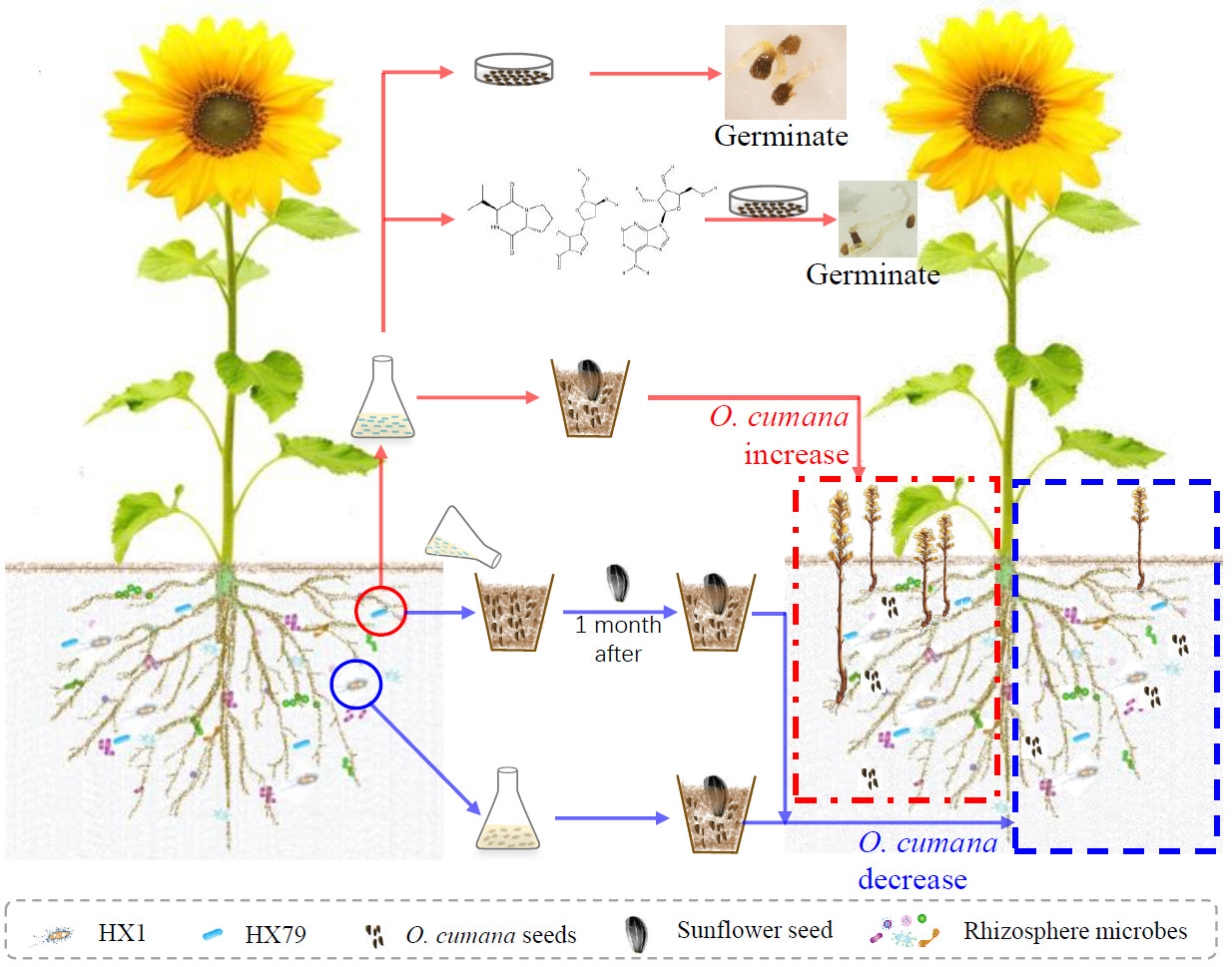
Microbial community roles and chemical mechanisms in the parasitic development of Orobanche cumana
- 13 June 2022
A comparative study of flow cytometry-sorted communities and shotgun viral metagenomics in a Singapore municipal wastewater treatment plant
- 28 July 2022
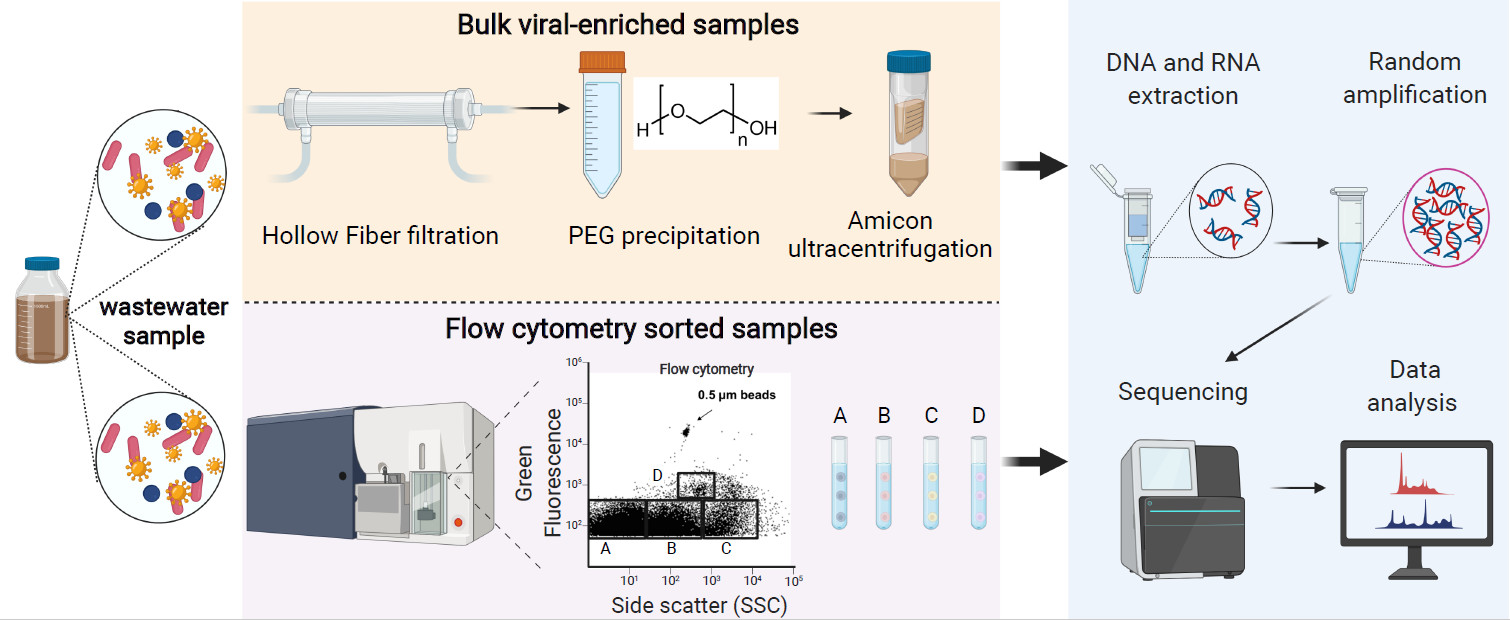
This study elucidates the practical protocol of applying fluorescence-activated cell sorting (FACS) and metagenomics on DNA and RNA viromes. This is the first systematic analysis of viral metagenomics in a municipal wastewater treatment plant with the Modified Ludzack–Ettinger (MLE) process. Coupling FACS with metagenomics improved the detection and genome assembly of certain rare viral species. In conclusion, bulk viral metagenomics and FACS-coupled metagenomics delivered complementary results.
StrainPanDA: Linked reconstruction of strain composition and gene content profiles via pangenome-based decomposition of metagenomic data
- 01 August 2022
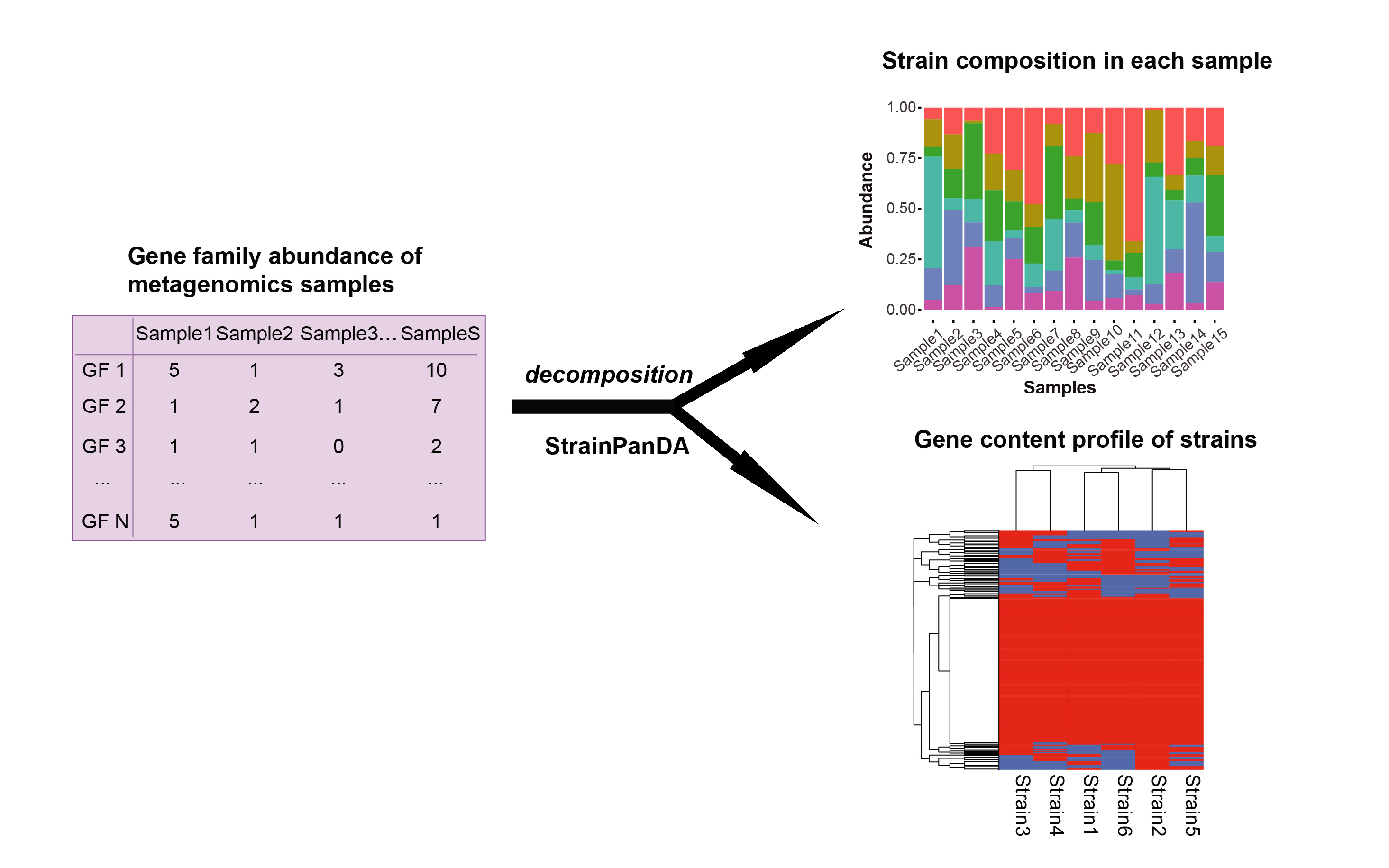
Strain-level Pangenome Decomposition Analysis (StrainPanDA) uses the pangenome coverage profile of multiple metagenomic samples to simultaneously reconstruct the composition and gene content variation of coexisting strains in microbial communities. StrainPanDA allows accurate and robust inference of strain composition and gene content profiles on synthetic data sets. Linked reconstruction of strain composition and gene content profiles provided by StrainPanDA furthers our understanding of the relationship between microbial adaptation and strain-specific functions (e.g., nutrient utilization and pathogenicity).
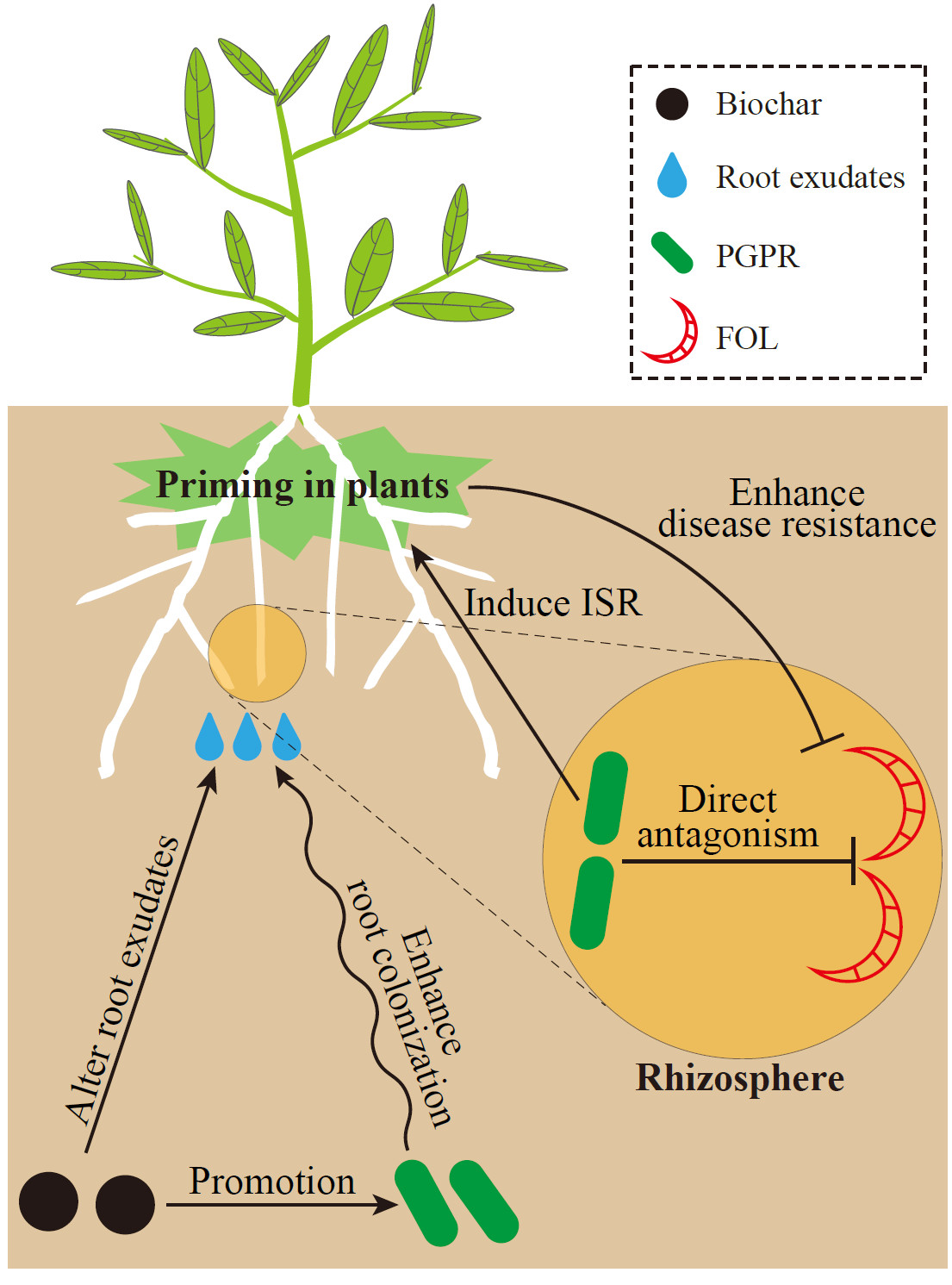
Tomato microbiome under long-term organic and conventional farming
- 19 August 2022
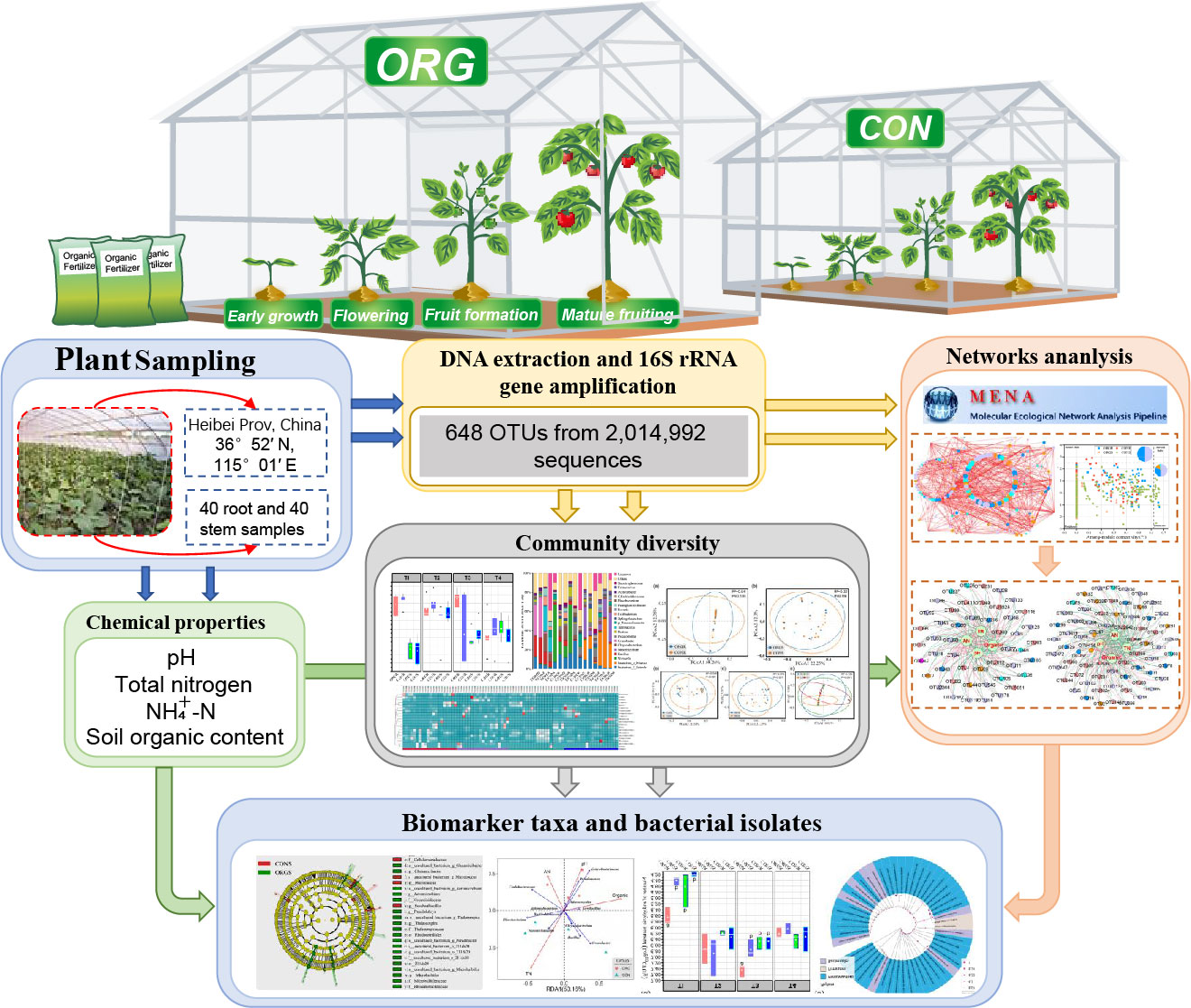
The compartment niche is the main reason behind the shifts in endophytic bacterial communities. Long-term organic greenhouse exerted limited influence on the variations of endophytic bacterial communities. Organic greenhouse and root had more complex co-occurrence networks than conventional greenhouse and stem, respectively. Cultivable method results found that Protecbacteria, Bacteriodes, and Actinobacteria are the dominant phyla in the endophytes.
Sangerbox: A comprehensive, interaction-friendly clinical bioinformatics analysis platform
- 08 July 2022
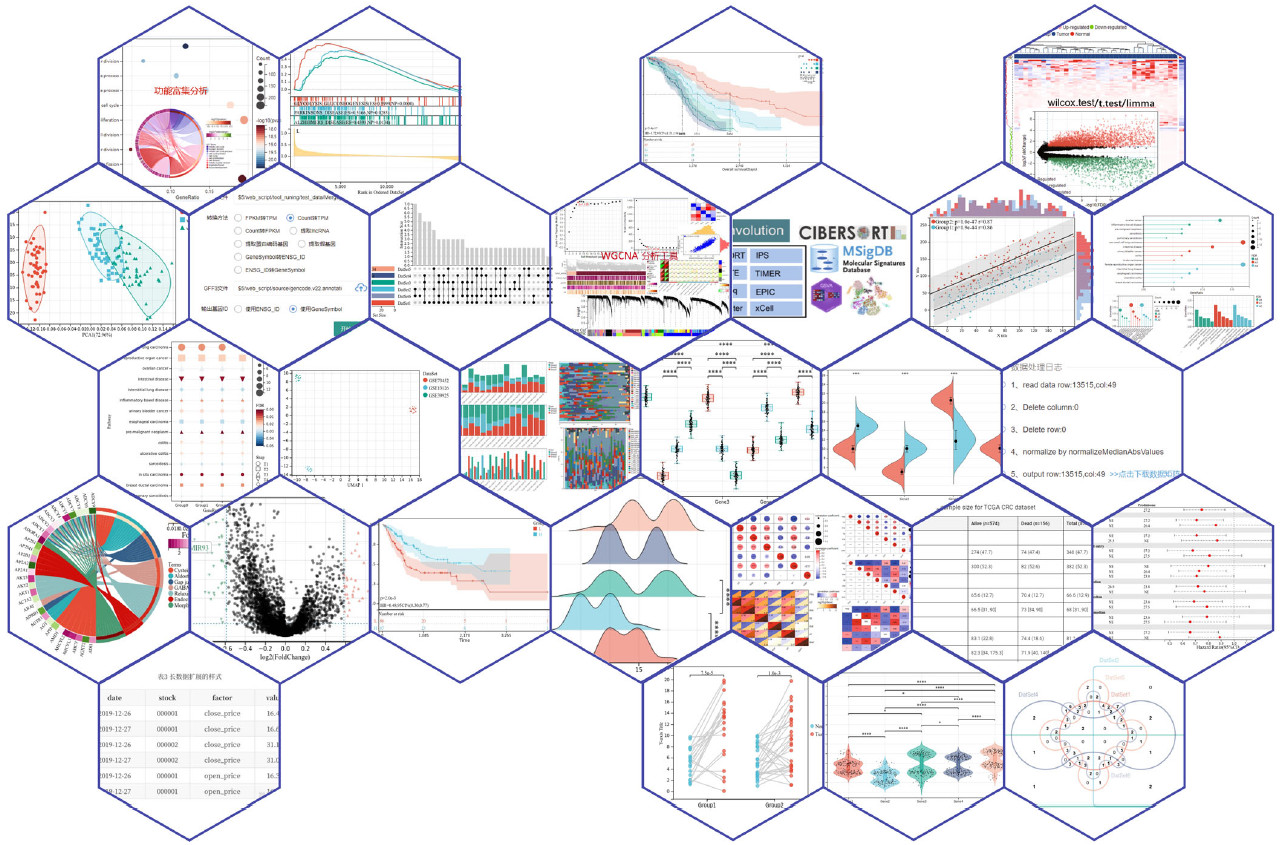

Sangerbox with a user-friendly interface supports differential analysis, correlation analyses, pathway enrichment analysis, weighted correlation network analysis, and so on. A new interactive plotting system that allows users to adjust the parameters in the image. It has organized GEO, TCGA, ICGC, and other databases; a rapid batch processing reduces the difficulty in data acquirement, greatly improving the efficiency.
Impact of tobacco smoking on oral cancer genetics—A next-generation sequencing perspective
- 04 August 2022
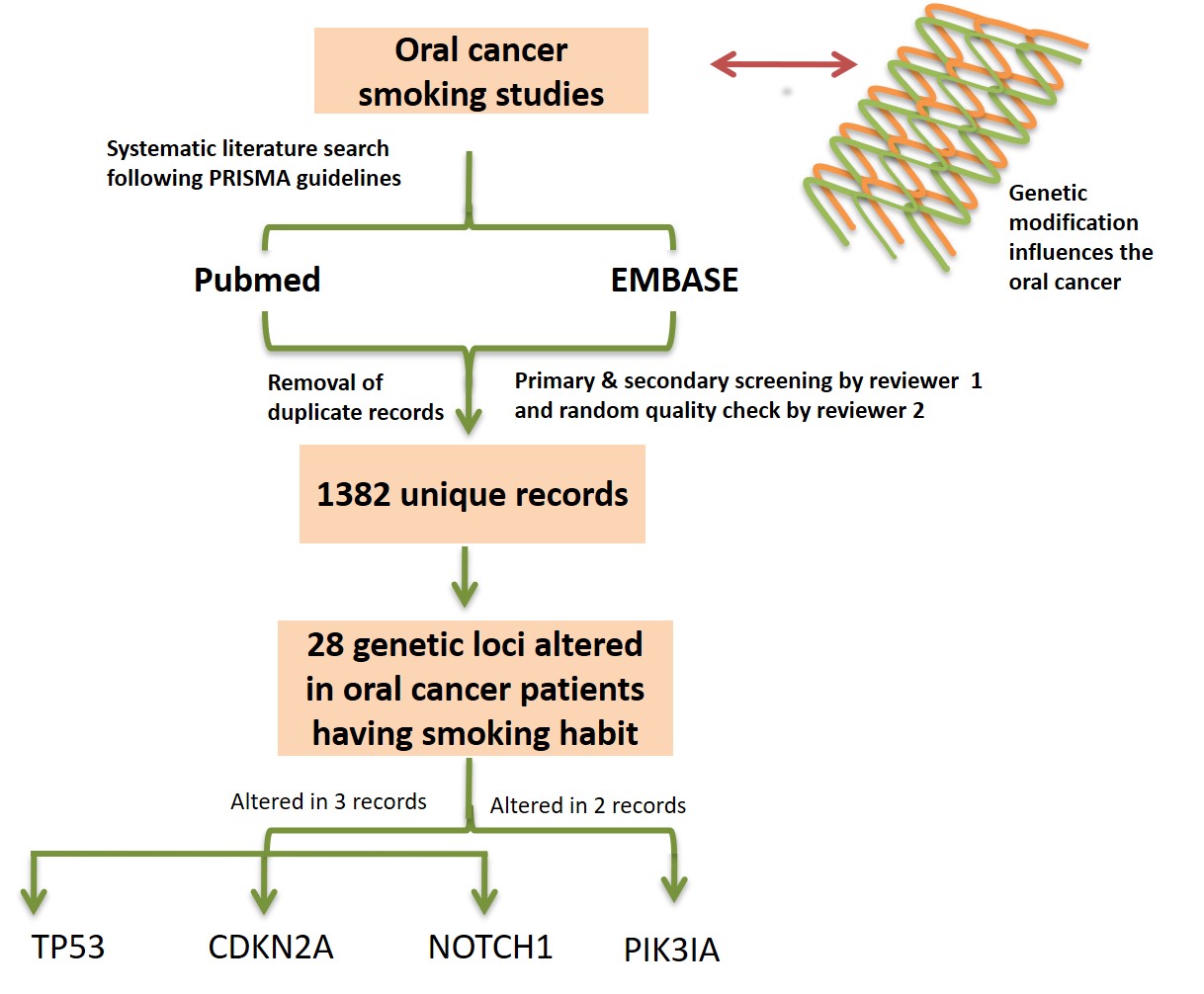
This study identified a total of 28 genetic loci (comprising 31 genes), which were found to be altered in oral cancer patients having a habit of tobacco smoking. Three loci, that is, 17p13.1 (TP53), 9p21.3 (CDKN2A), and 9q34.3 (NOTCH1) were found to be modified and common in three records whereas one locus, that is, 3q26.32 (PIK3CA) was found to be modified and common in two records. This study suggests that oral cancer patients should be categorized into different subgroups based on (i) genetic signatures, and (ii) smoking status, then the treatment strategies for each group should be precisely defined.