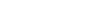Beyond personal space: Unveiling the transmission pattern of the human gut and oral microbiome
- 22 March 2023
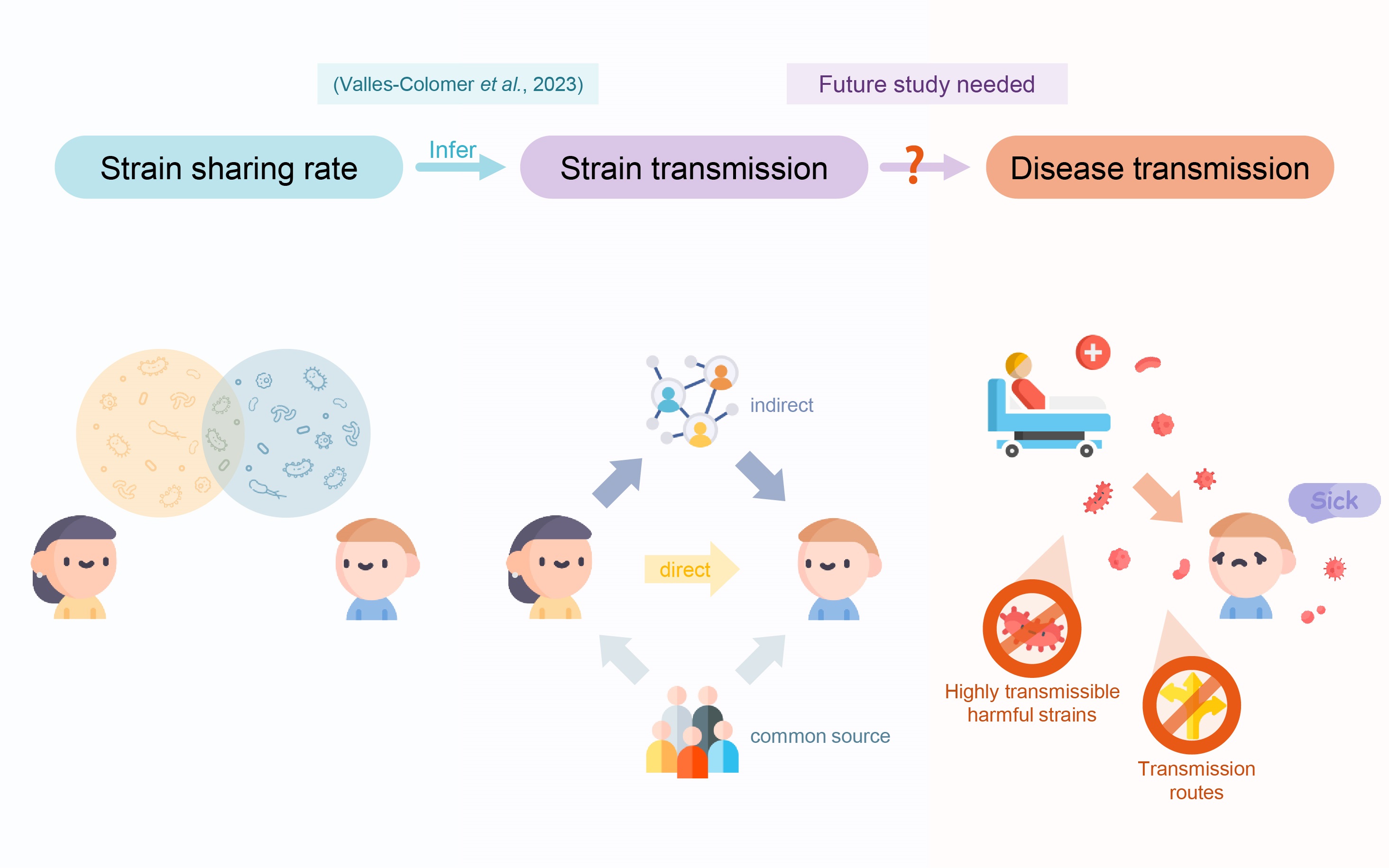
Valles-Colomer et al. evaluated the rate of microbial strain sharing to infer person-to-person microbial transmission. The presence of shared strains does not necessarily indicate direct transmission between individuals. Future research should investigate the potential for noncommunicable diseases to be transmitted through microbial strain transmission. Identification of harmful strains with high transmissibility and transmission routes can enable the development of targeted interventions.
Integration of multiomics with precision nutrition for gestational diabetes: Study protocol for the Westlake Precision Birth Cohort
- 06 February 2023
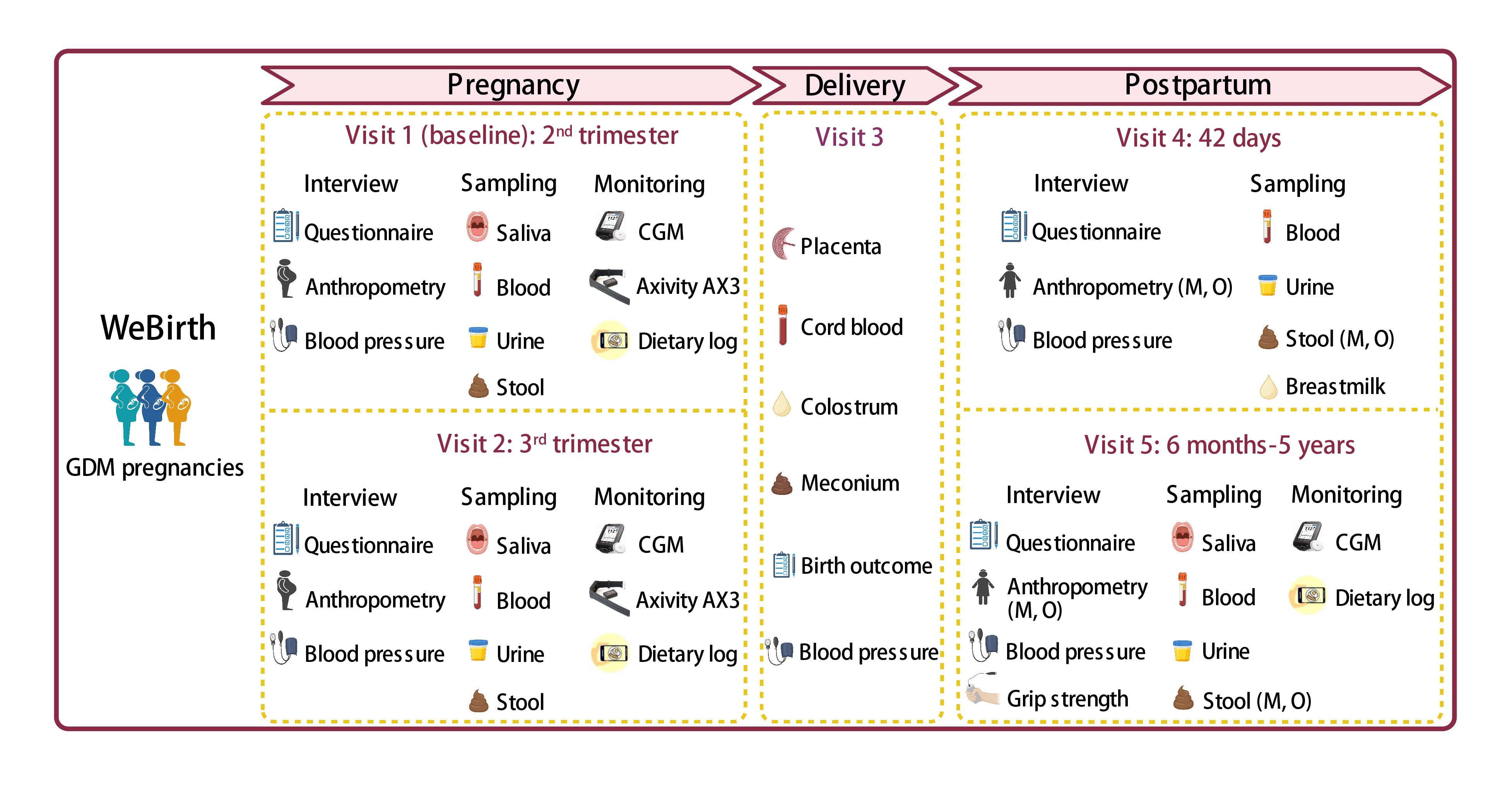
We established a prospective birth cohort, the Westlake Precision Birth Cohort (WeBirth), based on 2000 pregnant women with gestational diabetes mellitus (GDM) in the second trimester and their offspring. The WeBirth provides a new framework for prospective birth cohort study with sophisticated integration of precision nutrition, wearable devices, and multiomics data collection among patients with GDM.
Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp
- 08 May 2023
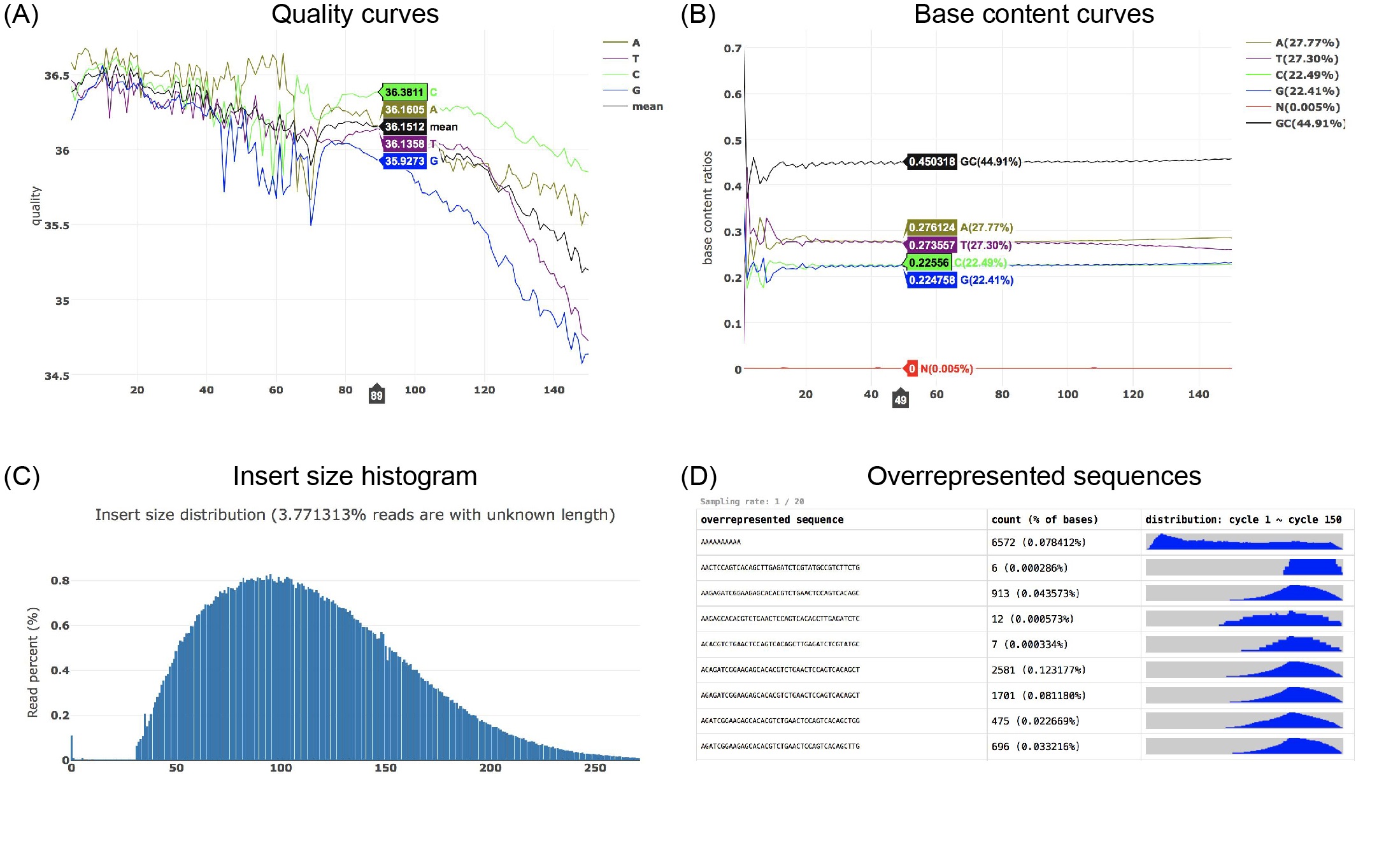
Fastp is a widely adopted tool for FASTQ data preprocessing and quality control. It is ultrafast and versatile and can perform adapter removal, global or quality trimming, read filtering, unique molecular identifier processing, base correction, and many other actions within a single pass of data scanning. Fastp has been reconstructed and upgraded with some new features. Compared to fastp 0.20.0, the new fastp 0.23.2 is even 80% faster.
Quantitative microbiome profiling reveals the developmental trajectory of the chicken gut microbiota and its connection to host metabolism
- 25 April 2023
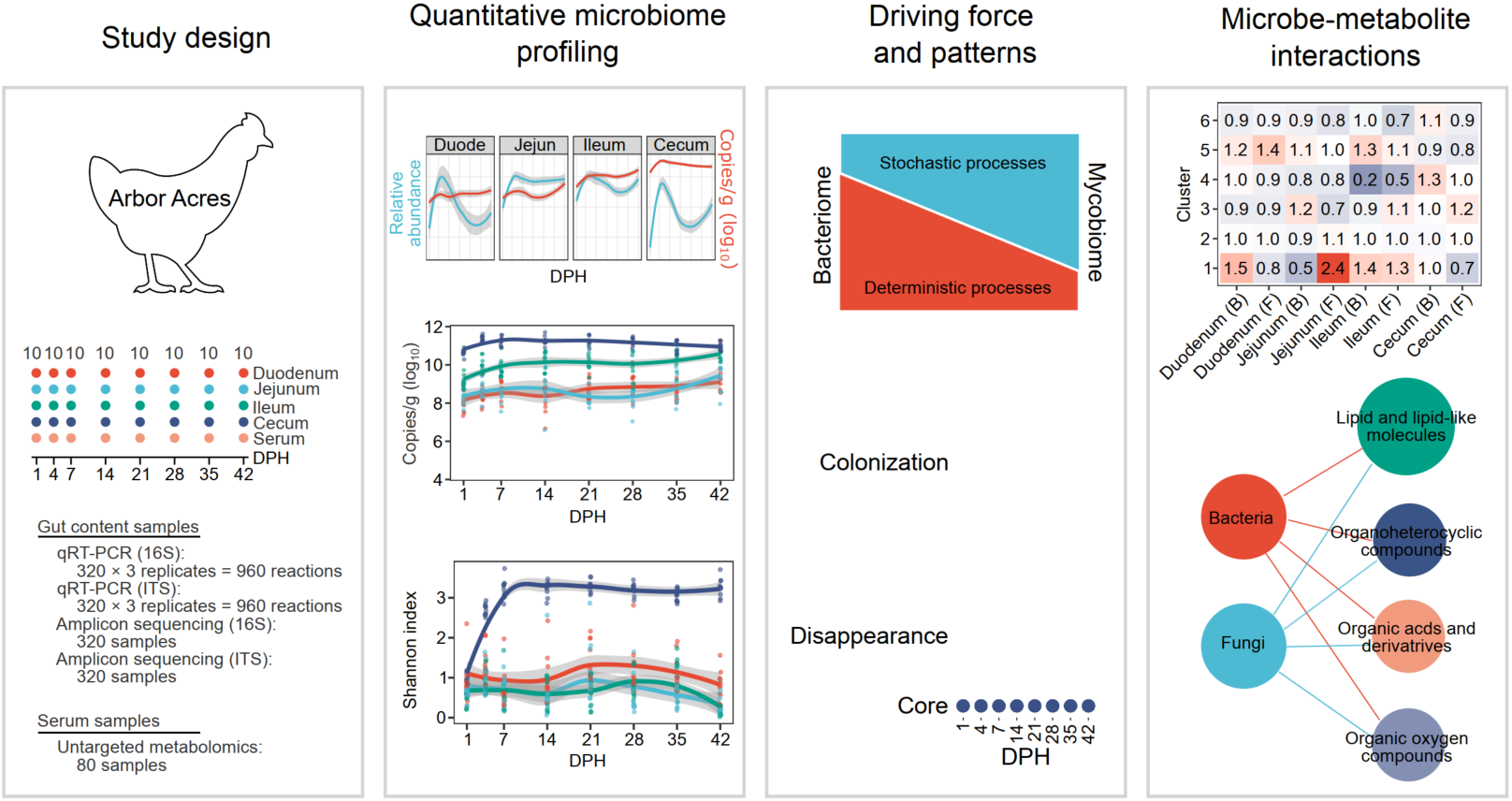
In this study, we revealed the developmental trajectory of the broiler chicken gut bacteriome and mycobiome by combining high-throughput sequencing with a microbial load quantification assay. Furthermore, we provided links between the absolute changes in chicken gut microbiota and their serum metabolite variations. The present study provided a full landscape of chicken gut microbiota development in a quantitative manner, and the associations between gut microbes and chicken serum metabolites further highlight the impact of gut microbiota in chicken growth and development.
Immunomagnetic-bead enriched culturomics (IMBEC) for isolating pathobionts from feces of colorectal cancer patients
- 05 April 2023
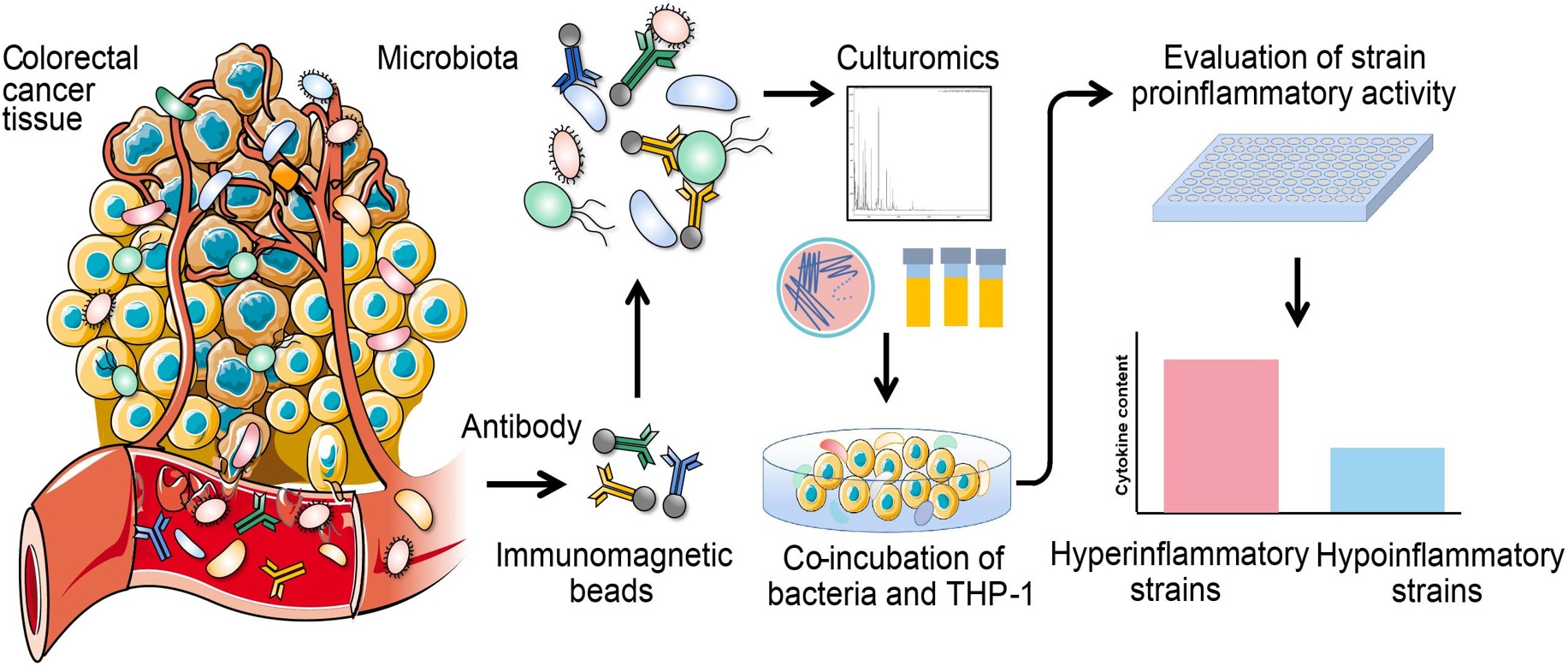
We developed a novel method, immunomagnetic bead-enriched culturomics (IMBEC), which can obtain potential pathobionts from the precultivated colorectal cancer (CRC) fecal samples. We optimized the IMBEC process from the perspective of coupling reagents and magnetic beads washing lotion. Not only in CRC but it has also been reported that antibodies produced by pathobiont strains exist in the blood of patients with other diseases, so the method of precise isolation of disease-related strains by IMBEC is expected to be a powerful method to accurately isolate pathobiont strains.
Intratumoral microbiota is associated with prognosis in patients with adrenocortical carcinoma
- 05 April 2023
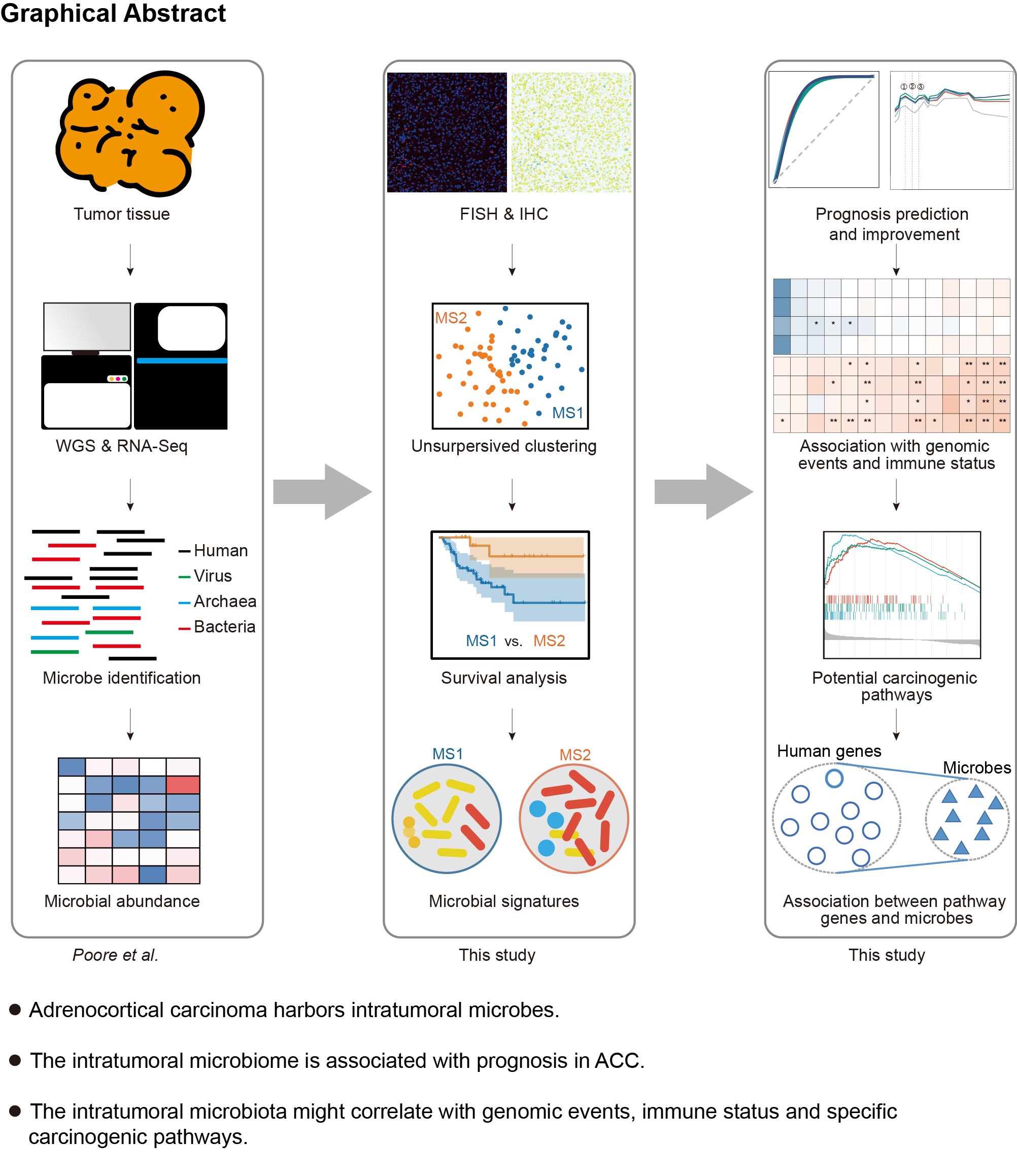
The Cancer Genome Atlas primary tumors were subject to whole-genome sequencing and RNA-Seq, in which microbial abundance (comprising virus, archaea, and bacteria) was identified and normalized by Poore et al. (left panel). To confirm the presence of intratumor bacteria, 37 adrenocortical carcinoma tissue microarray chips from the in-house cohort were stained for fluorescence in situ hybridization and immunohistochemistry examination. Unsupervised clustering was performed on microbiome data to explore natural clusters of patients with distinct microbiomes, thereby influencing prognoses. The microbial signatures associated with prognosis were further determined (middle panel). Defined microbial signatures were tested to improve prognosis prediction. The intratumoral microbiome was found to be associated with immune status, genetic events, and molecular pathways (right panel).
shinyCircos-V2.0: Leveraging the creation of Circos plot with enhanced usability and advanced features
- 08 May 2023
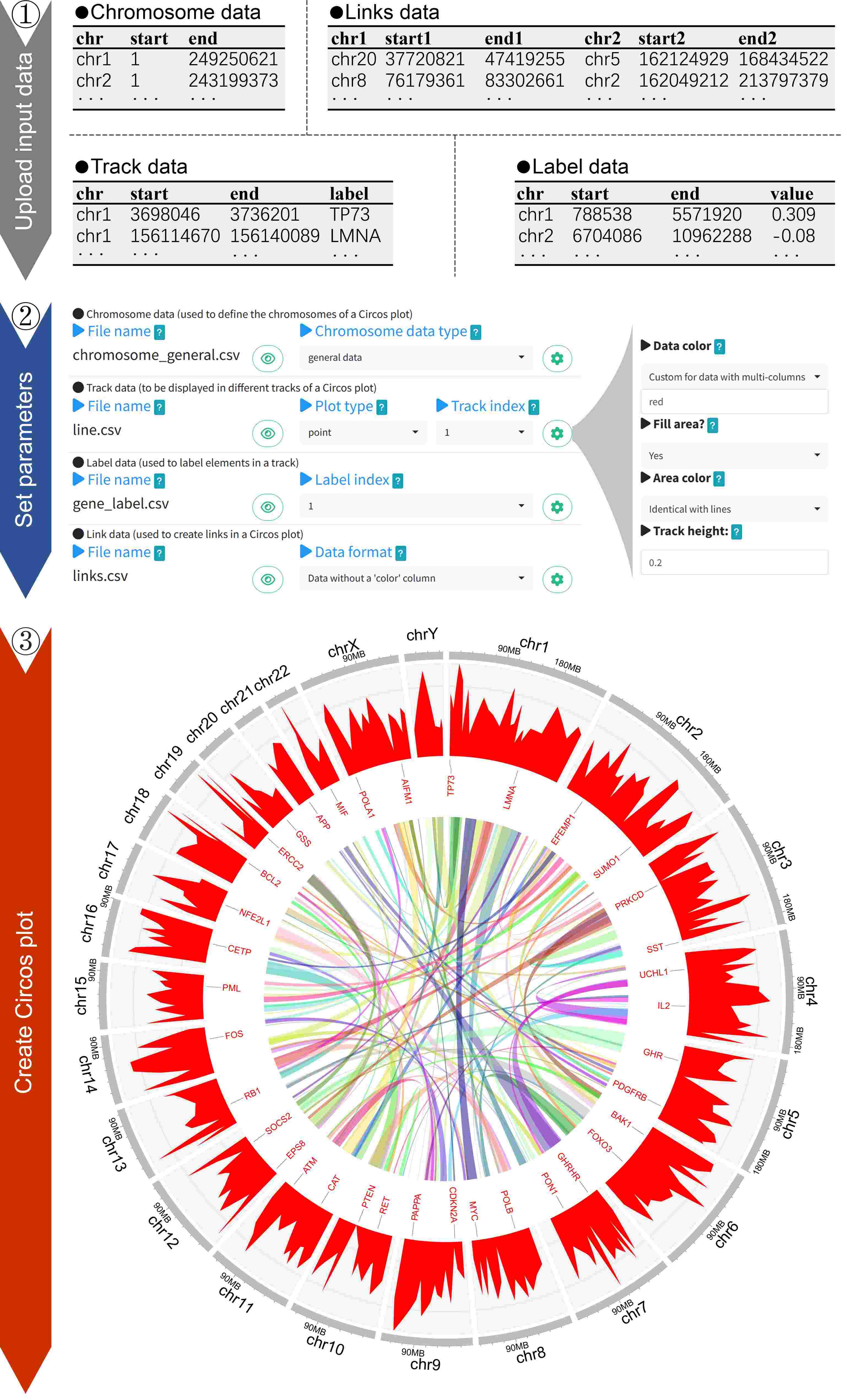
We developed a web application called shinyCircos-V2.0 for creating Circos plots. The process of creating a Circos plot with shinyCircos-V2.0 involves three simple steps: uploading the input data, setting the plotting parameters, and generating the Circos plot. Our application offers a wide range of plotting parameters that can be used to enhance the appearance of the resulting Circos plot. Overall, shinyCircos-V2.0 simplifies the creation of Circos plots and provides researchers with a powerful tool for data visualization.
Ontology characterization, enrichment analysis, and similarity calculation-based evaluation of disease–syndrome–formula associations by applying SoFDA
- 10 January 2023
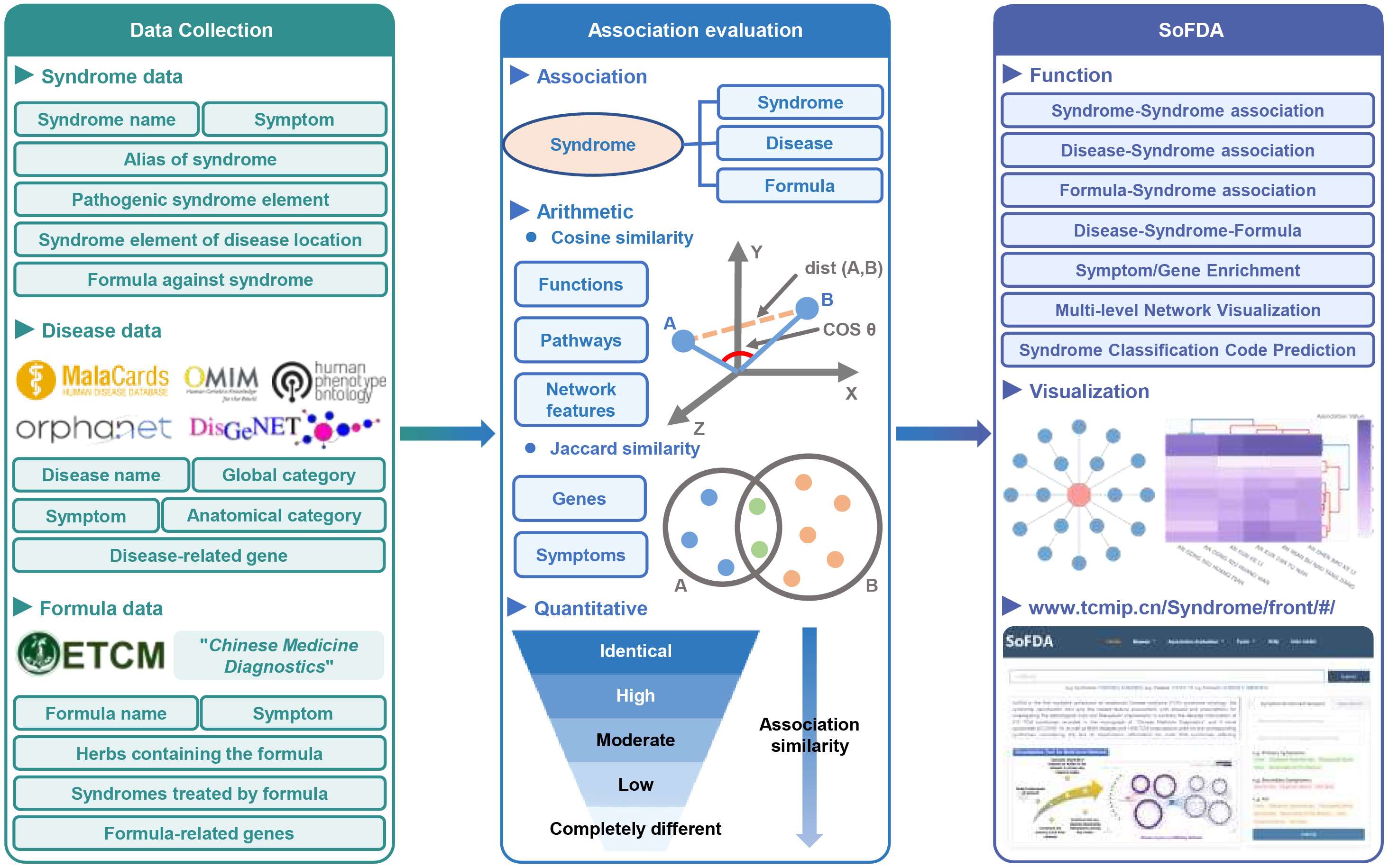
SoFDA is the first integrated web platform, SoFDA (http://www.tcmip.cn/Syndrome/front/), with a curated ontology of 319 traditional Chinese medicine syndromes and their relationships with genes, diseases, and formulas. This platform enables biomedical and pharmaceutical scientists to rank and filter the most promising associations for disease diagnosis and tailored interventions.
Schizophrenia and obesity: May the gut microbiota serve as a link for the pathogenesis?
- 04 April 2023
Research advances in probiotic fermentation of Chinese herbal medicines
- 19 February 2023

This review summarizes the advantages of probiotic fermentation of Chinese herbal medicine (CHM), fermentation techniques, probiotic strains, and future development of CHM fermentation. Besides, synthetic biology would harness microbial cell factories to de novo produce large amounts of CHM bioactive natural products with low-cost, and it might be an opportunity to bring CHM into modern biomanufacturing.
Interactions between extracellular vesicles and microbiome in human diseases: New therapeutic opportunities
- 06 February 2023
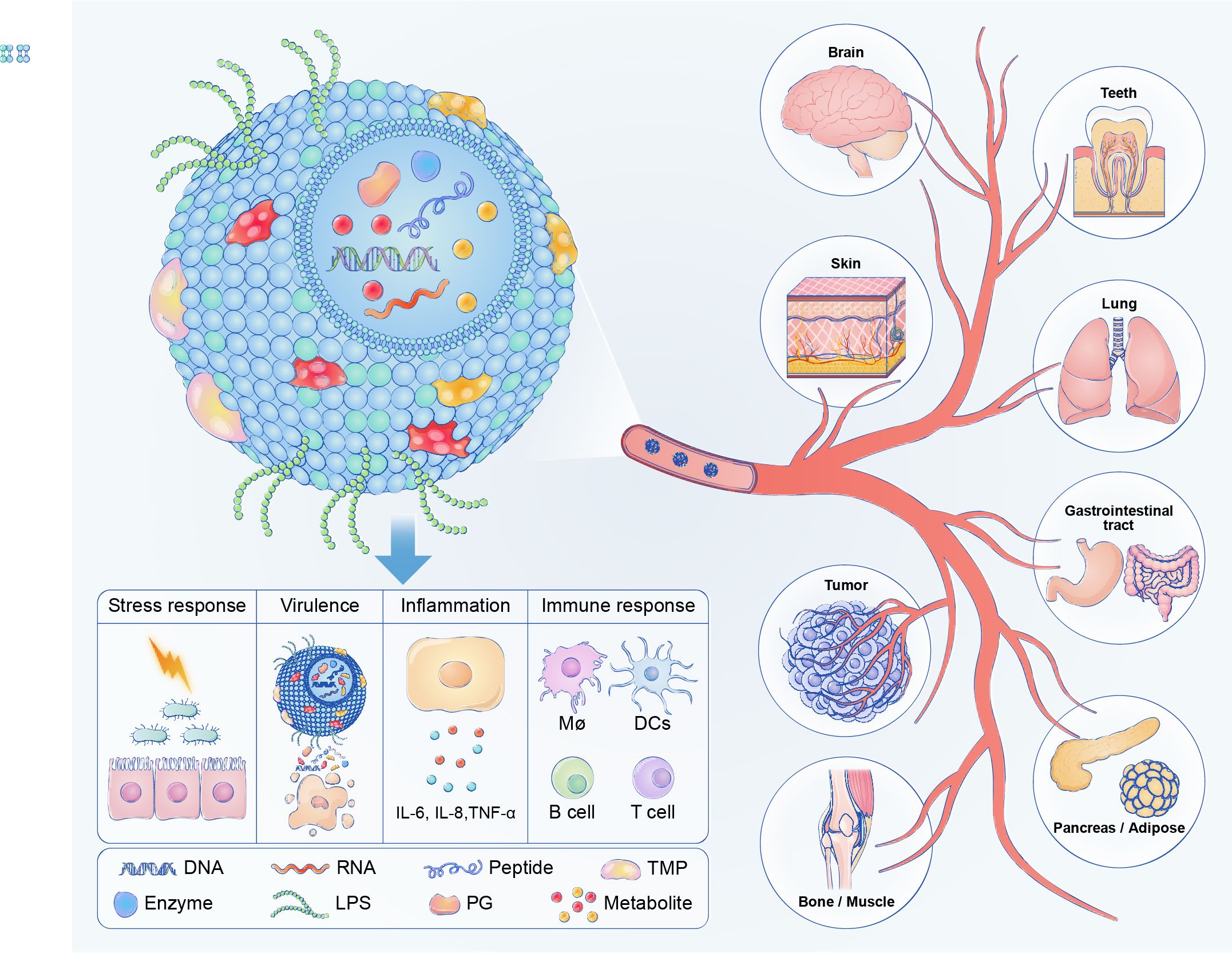
Bacterial extracellular vesicles (BEVs) derived from various commensal and pathogenic bacteria have been shown to participate in a wide range of physiological and pathological processes by transporting versatile bioactive molecules, including proteins, nucleic acids, lipids, and metabolites. BEVs are intensively involved in the onset and progression of various diseases and BEV-based platforms possess great potential in diagnostic analytics, vaccine design, and novel therapeutic development.
An overview of host-derived molecules that interact with gut microbiota
- 13 February 2023
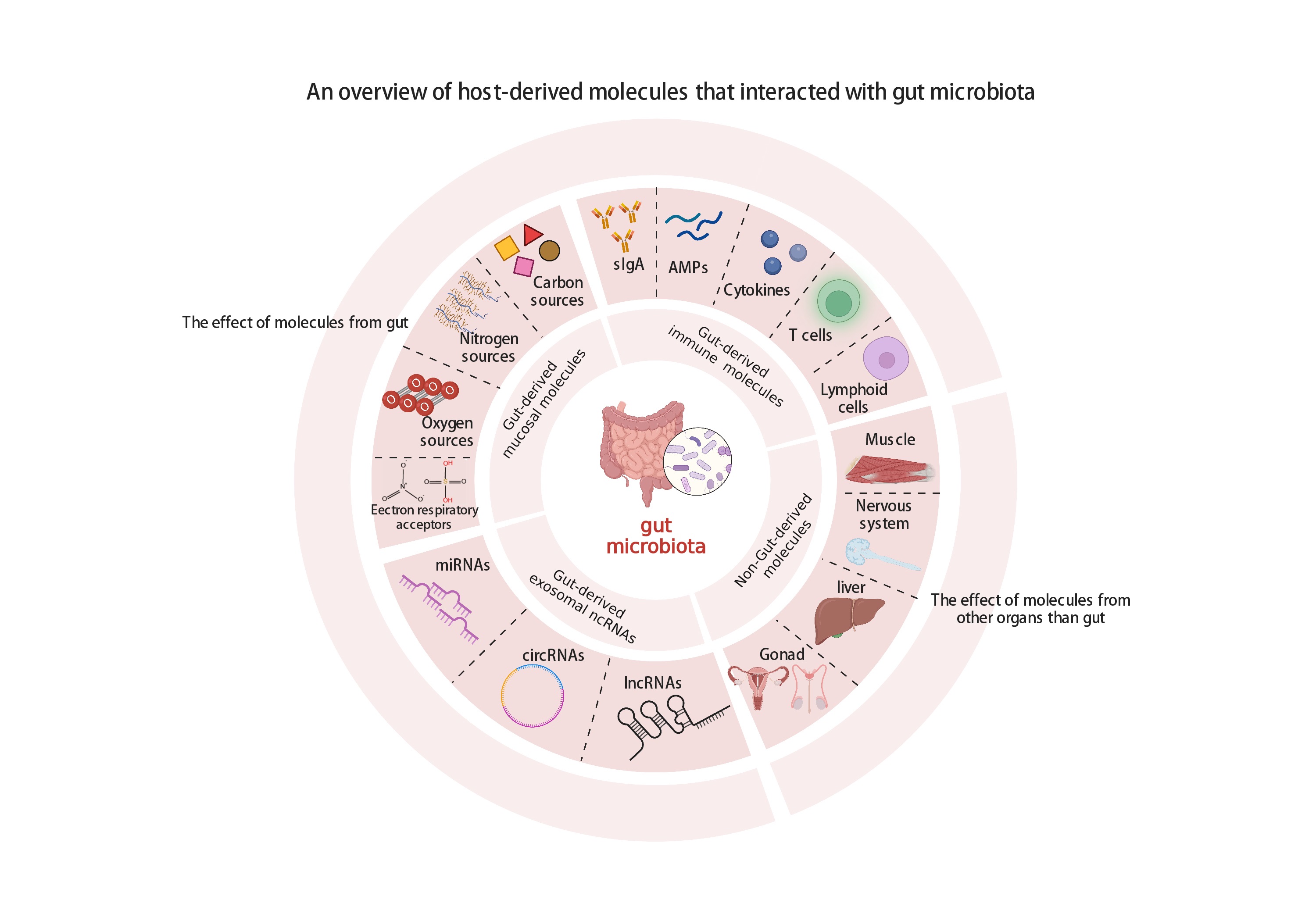
The mechanisms that mediate host regulation on gut microbiota, involved in the gut- and nongut-derived metabolites, including gut-derived immune system factors, sources related to gut-derived mucosal metabolites, gut-derived exosomes' noncoding RNA regulation, and nongut-derived metabolite-related sources are discussed to provide a systematic overview of how host metabolites govern the gut microbiota.
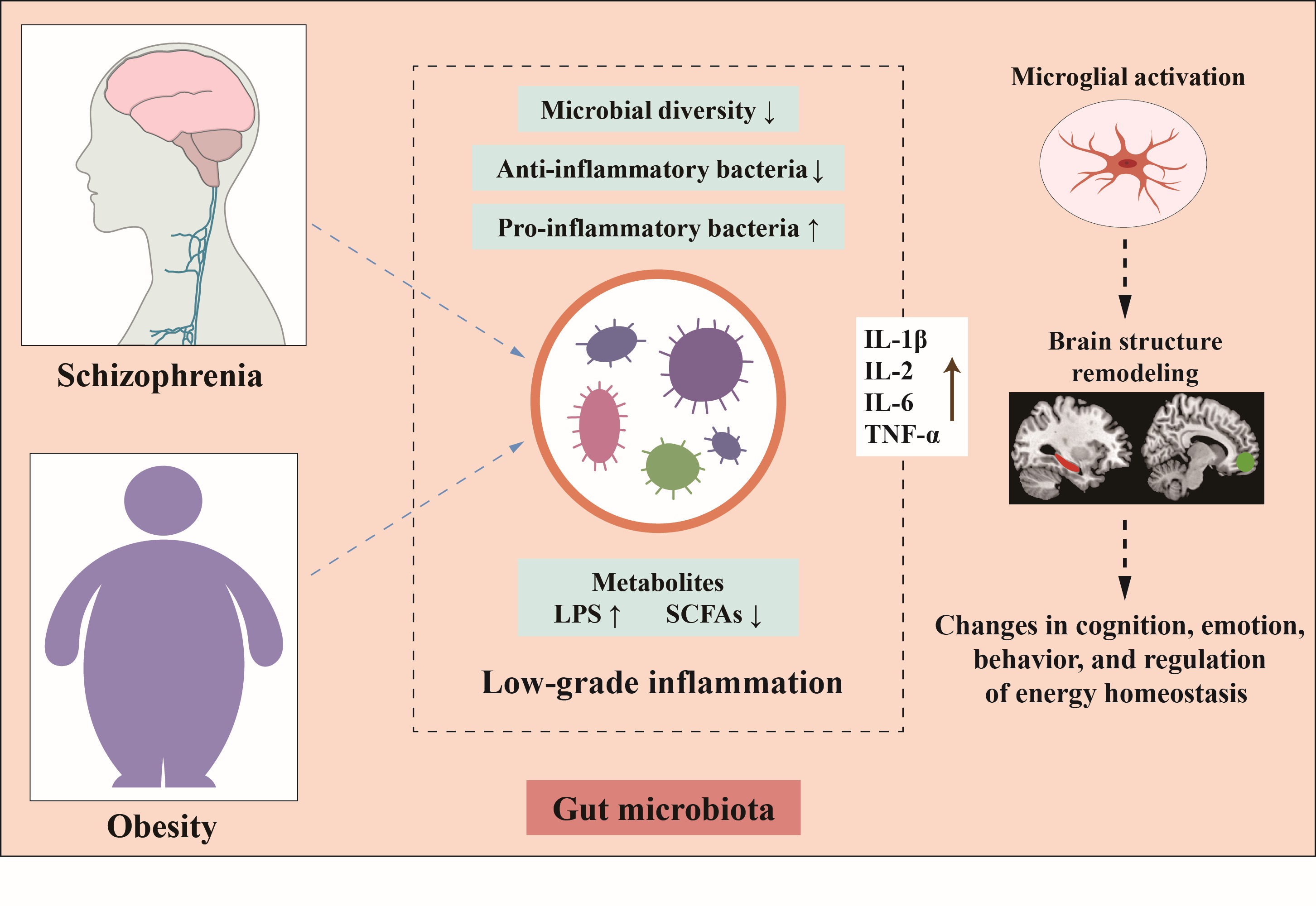
Plastisphere microbiome: Methodology, diversity, and functionality
- 31 March 2023
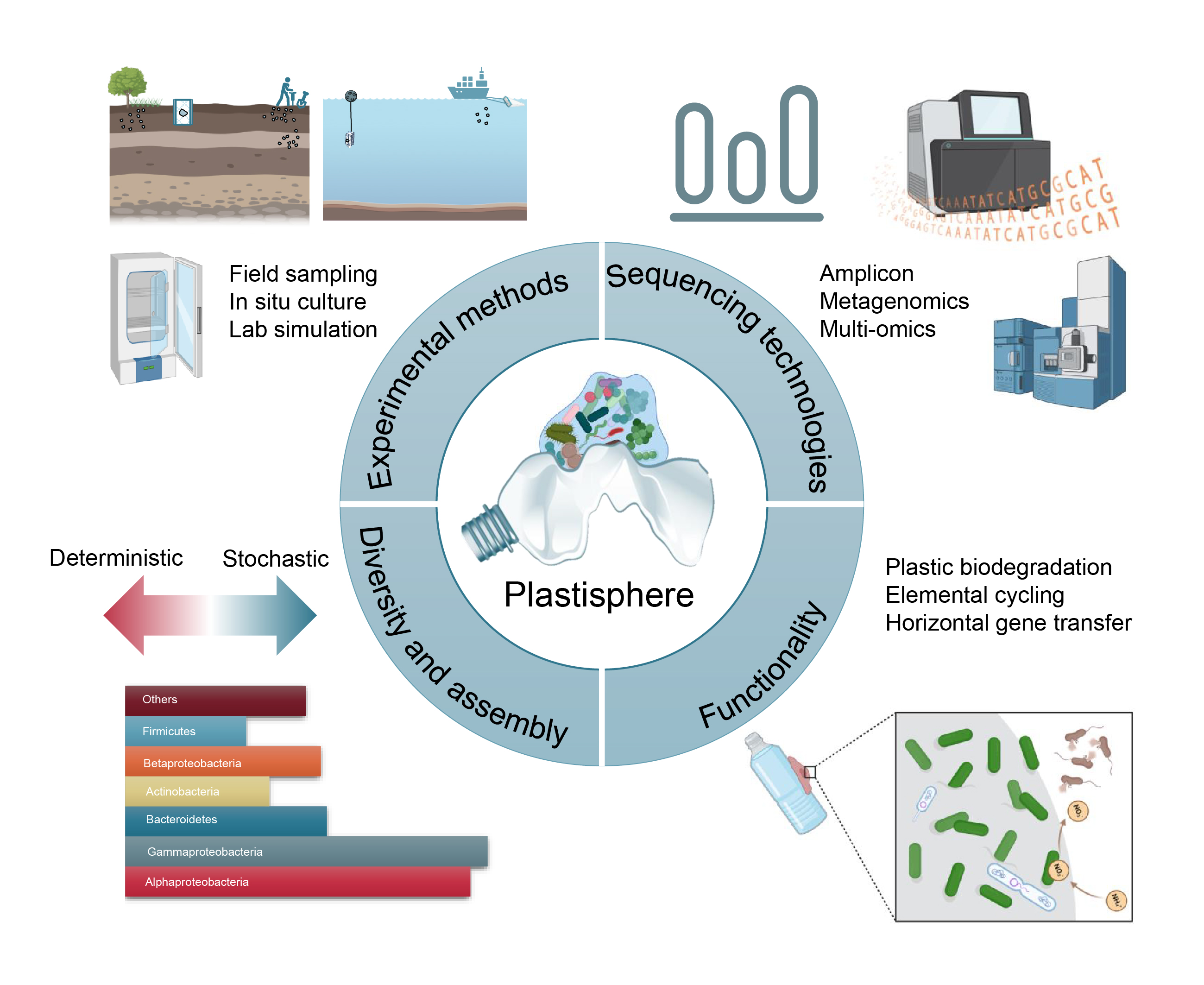
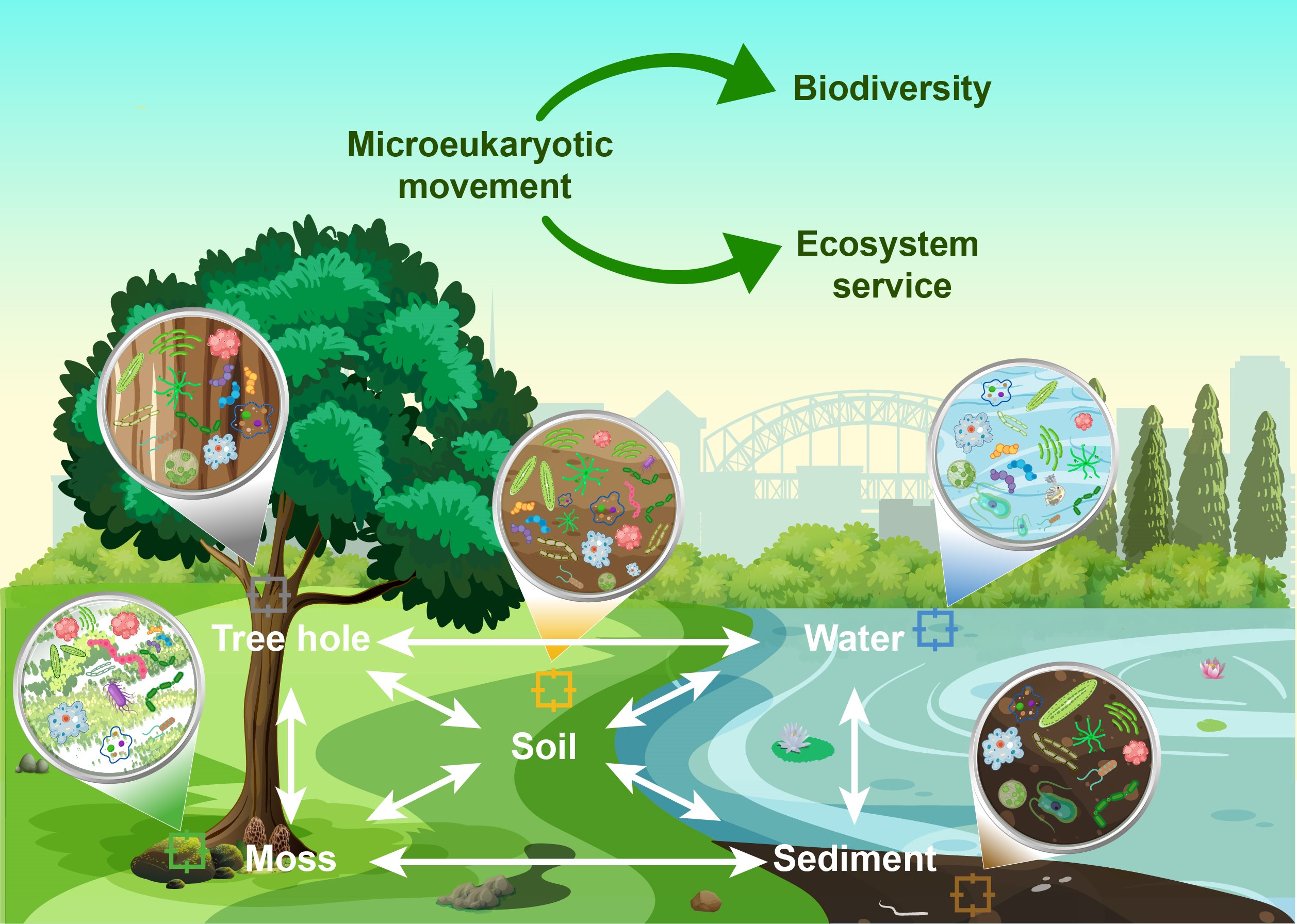
Linking biodiversity and ecological function through extensive microeukaryotic movement across different habitats in six urban parks
- 17 April 2023
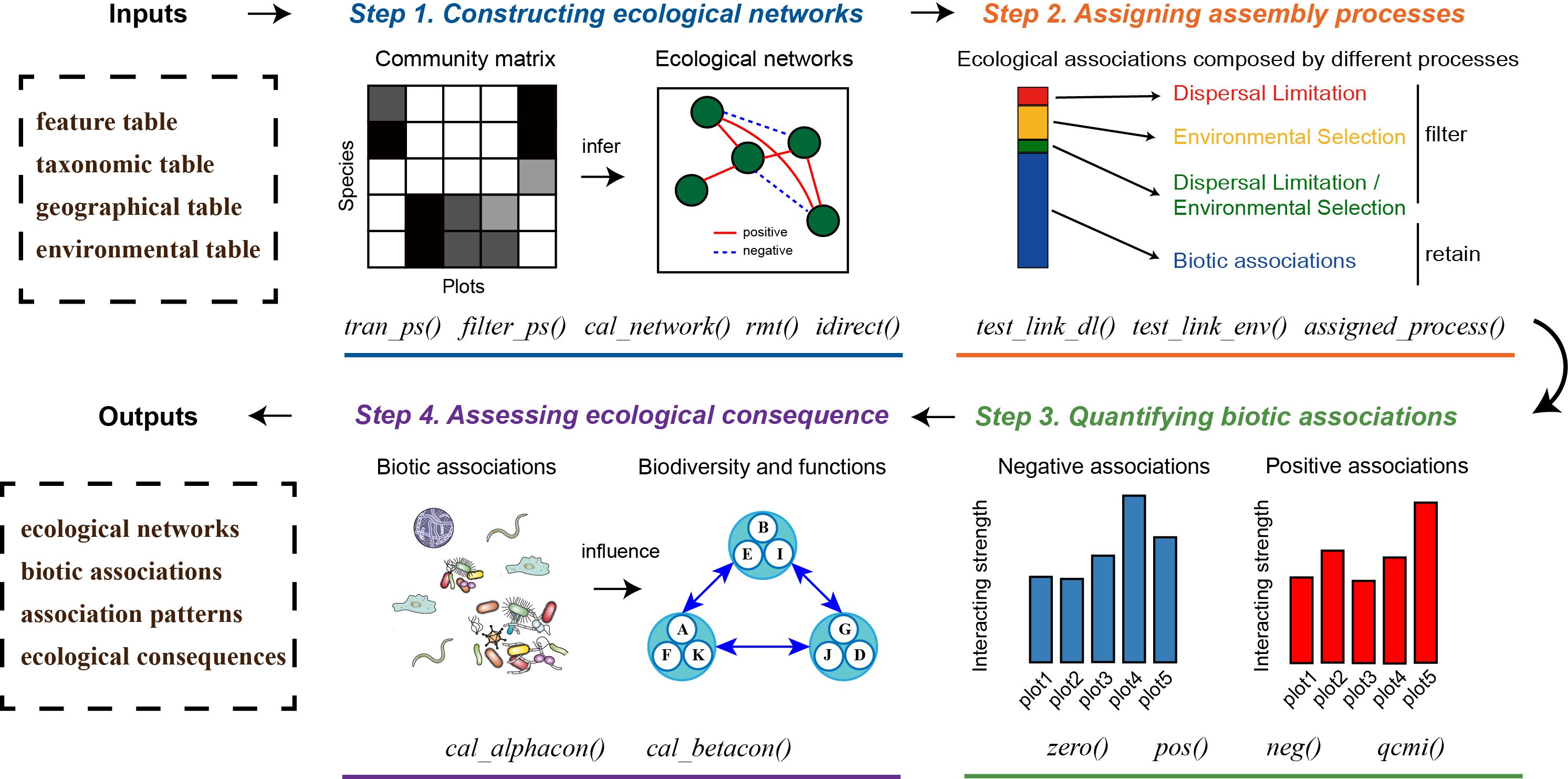
QCMI: A method for quantifying putative biotic associations of microbes at the community level
- 16 February 2023
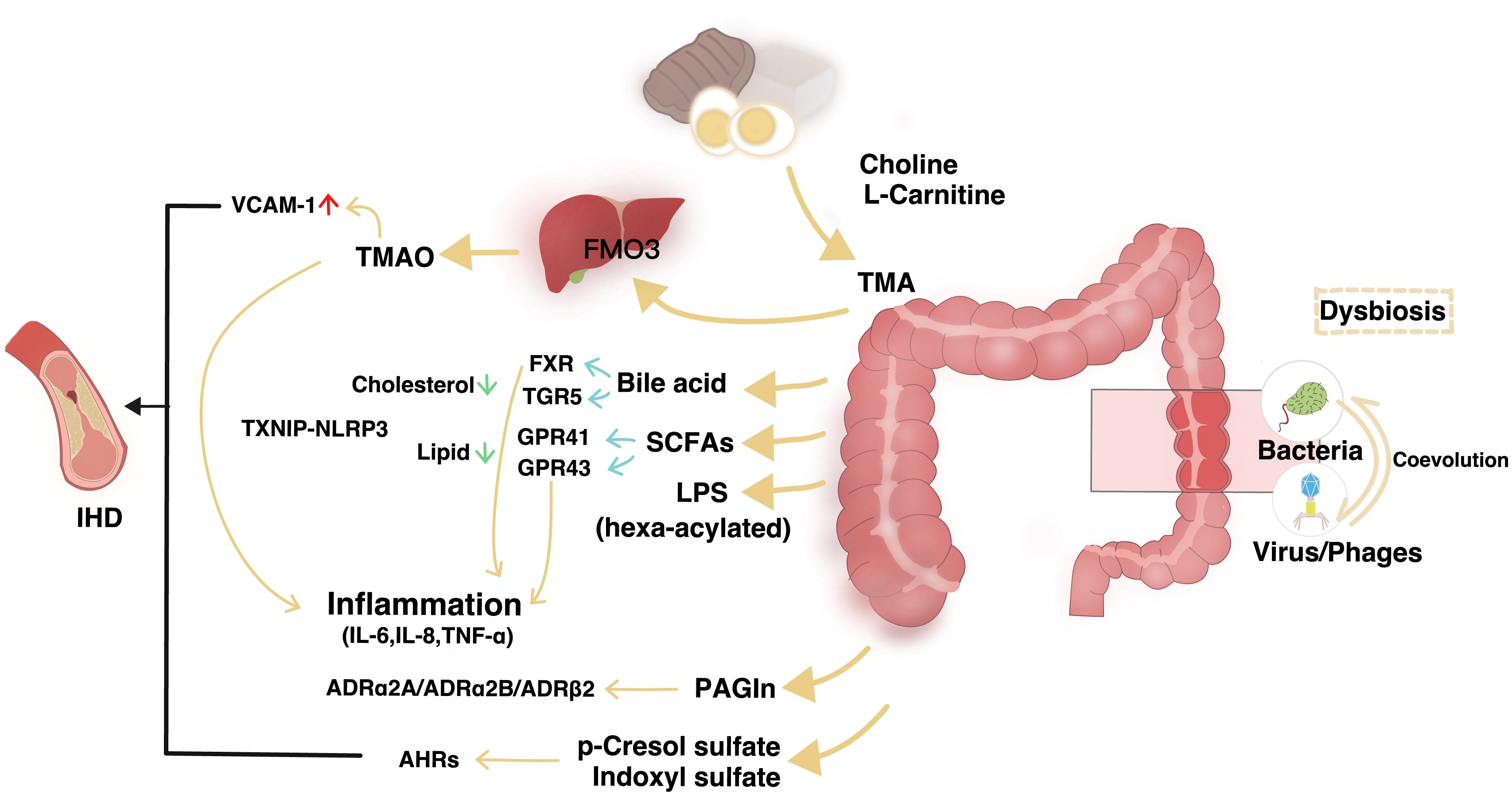
Microbiota-related metabolites fueling the understanding of ischemic heart disease
- 26 February 2023
Misuse of reporter score in microbial enrichment analysis
- 23 February 2023
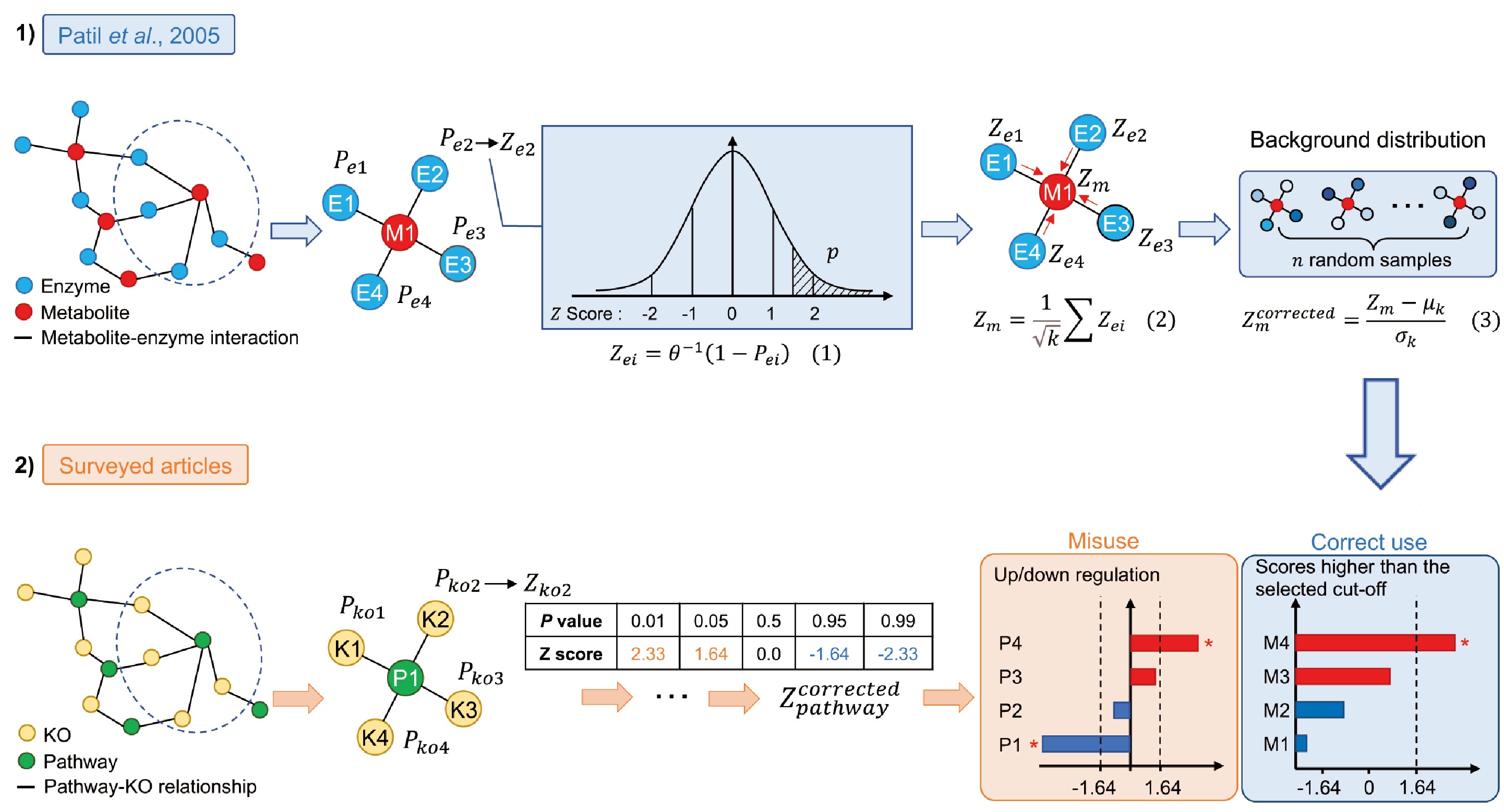
A modified new method for microbial enrichment analysis, reporter score was incorrectly used in many articles due to a lack of comprehensive and systematic understanding of the original method by the researchers, leading to a serious snowball effect. Here we describe the reasons for the misuse of reporter score and its negative impact on microbial research and hope this comment will facilitate community discussion on the importance of statistical rigor, informing future efforts to enhance reliable and reproducible research.
A high-salt diet induces synaptic loss and memory impairment via gut microbiota and butyrate in mice
- 21 March 2023
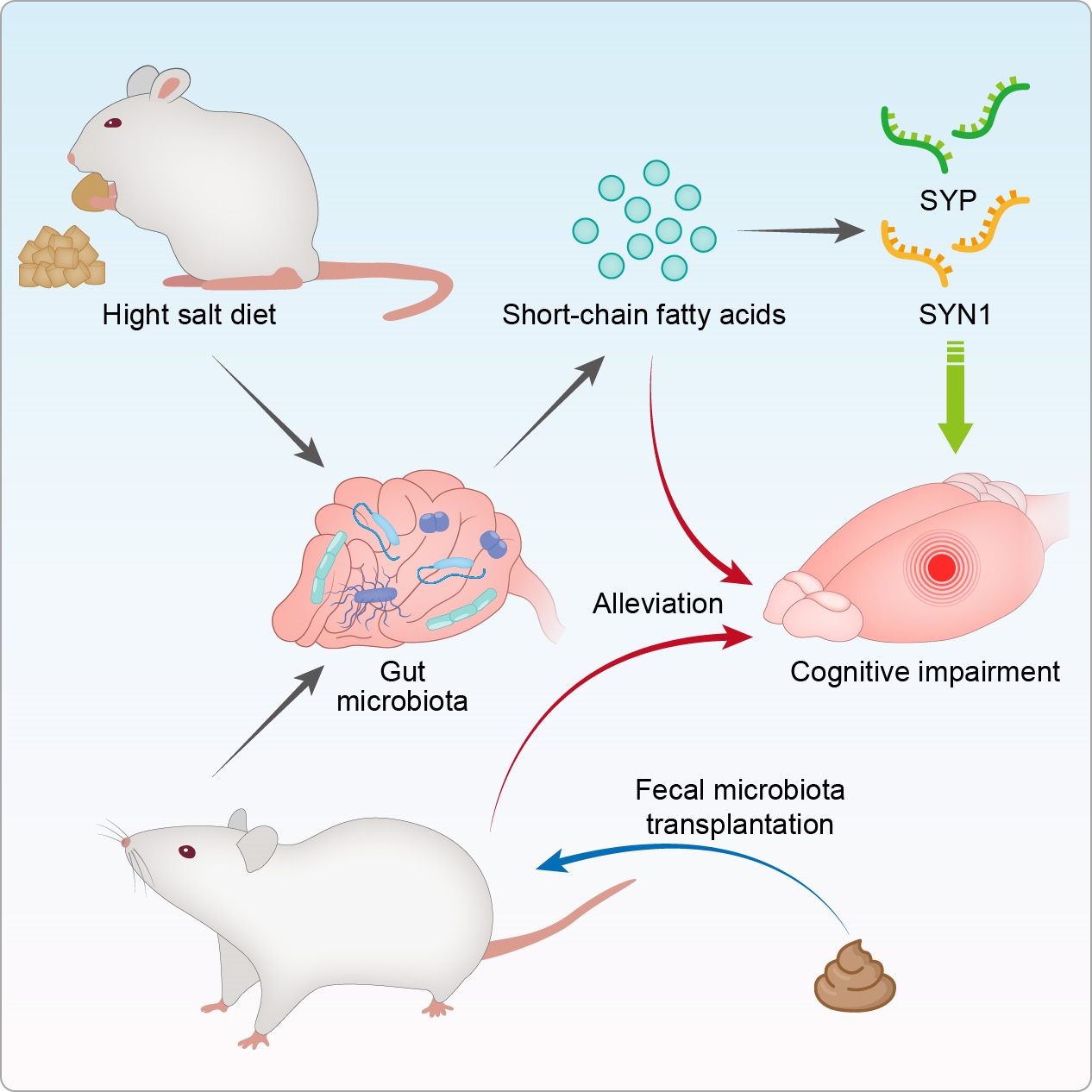
High-salt diet (HSD)-fed mice display cognitive impairment and lower synaptic proteins via changed gut microbiota composition and short-chain fatty acids production. Gut microbiota from HSD-fed mice impairs memory and synapse in normal salt diet-fed mice. Butyrate treatment partially reverses memory impairment in HSD-fed mice. Above all, this study indicates the important role of the gut microbiome and butyrate production in synaptic loss and memory impairment.
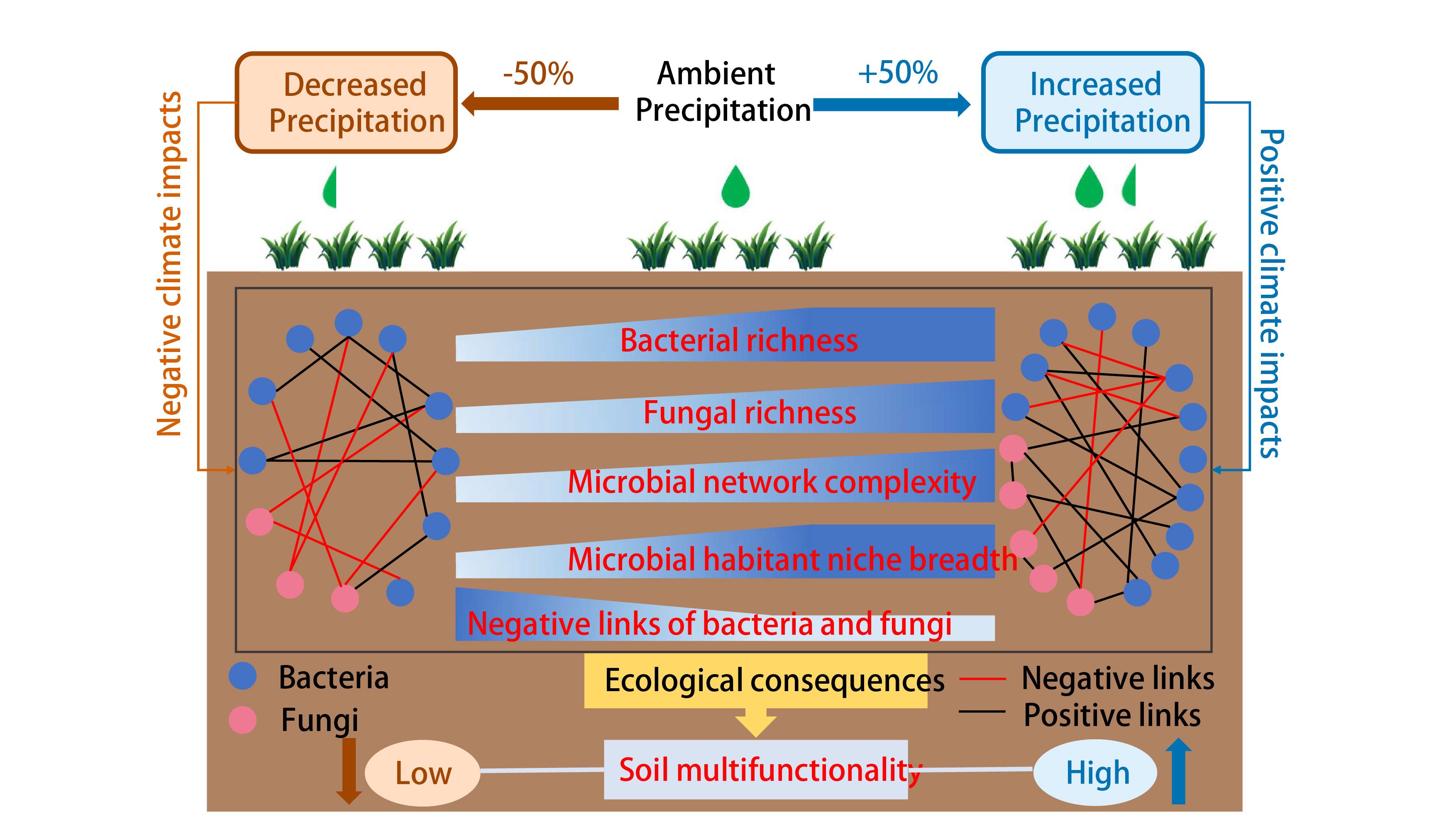
Decreased soil multifunctionality is associated with altered microbial network properties under precipitation reduction in a semiarid grassland
- 02 May 2023