iMeta Conference 2025: Creating high-impact international journals
- 15 October 2025

The iMeta Conference 2025, part of the iMeta Conference series, themed “Creating High-Impact International Journals,” held at the Huangjiahu Campus of Hubei University of Chinese Medicine from August 23rd to 25th, 2025, and focused on frontier topics such as microbiology, medicine, traditional Chinese medicine, botany, and research career development. The event aimed to support the development of researchers and strengthen the impact of academic journals. Through invited reports, thematic seminars, and poster presentations, the conference highlighted hot topics including multi-omics technologies, microbe-host interactions, AI-assisted research, live biotherapeutic products, and the modernization of traditional Chinese medicine. The event demonstrated the innovative momentum of interdisciplinary integration and technological convergence, providing an international platform for academic exchange and laying a foundation for building an innovative scientific research ecosystem and enhancing the global influence of Chinese academic journals.
High-throughput generic single-entity sequencing using droplet microfluidics
- 30 October 2025
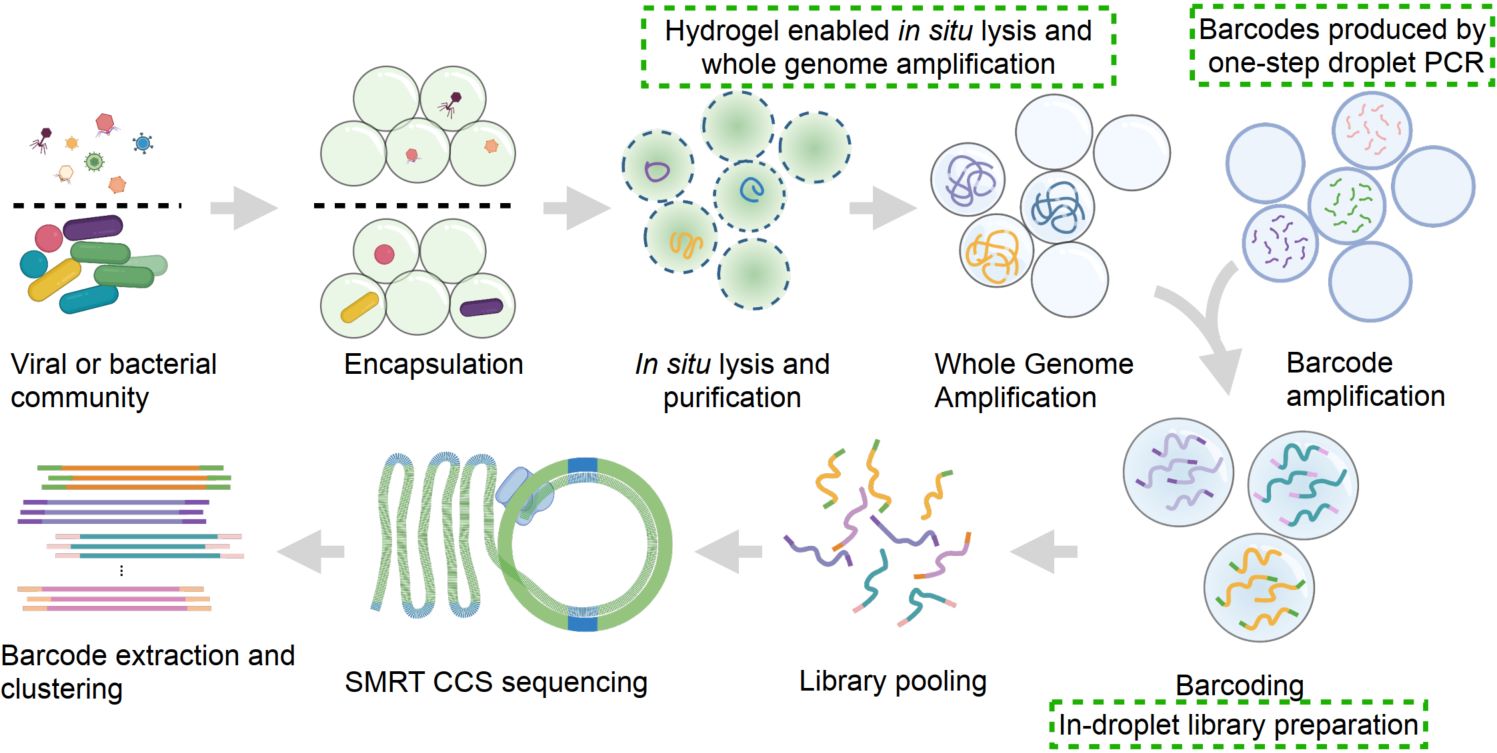
We present Generic Single Entity Sequencing (GSE-Seq), a droplet-based workflow that generates monoclonal barcodes by one-step PCR, performs dissolvable hydrogel-enabled reactions for genome processing, and attaches barcodes during in-droplet library preparation. Barcoded fragments are pooled and sequenced with PacBio long-read sequencing. Barcodes are extracted and clustered to recover single-entity genomes from viruses and bacteria, enabling high-throughput single-entity sequencing with long-read and accurate genome reconstruction.
Homoharringtonine suppresses acute myeloid leukemia progression by orchestrating EWSR1 phase separation in an m6A-YTHDF2-dependent mechanism
- 30 October 2025
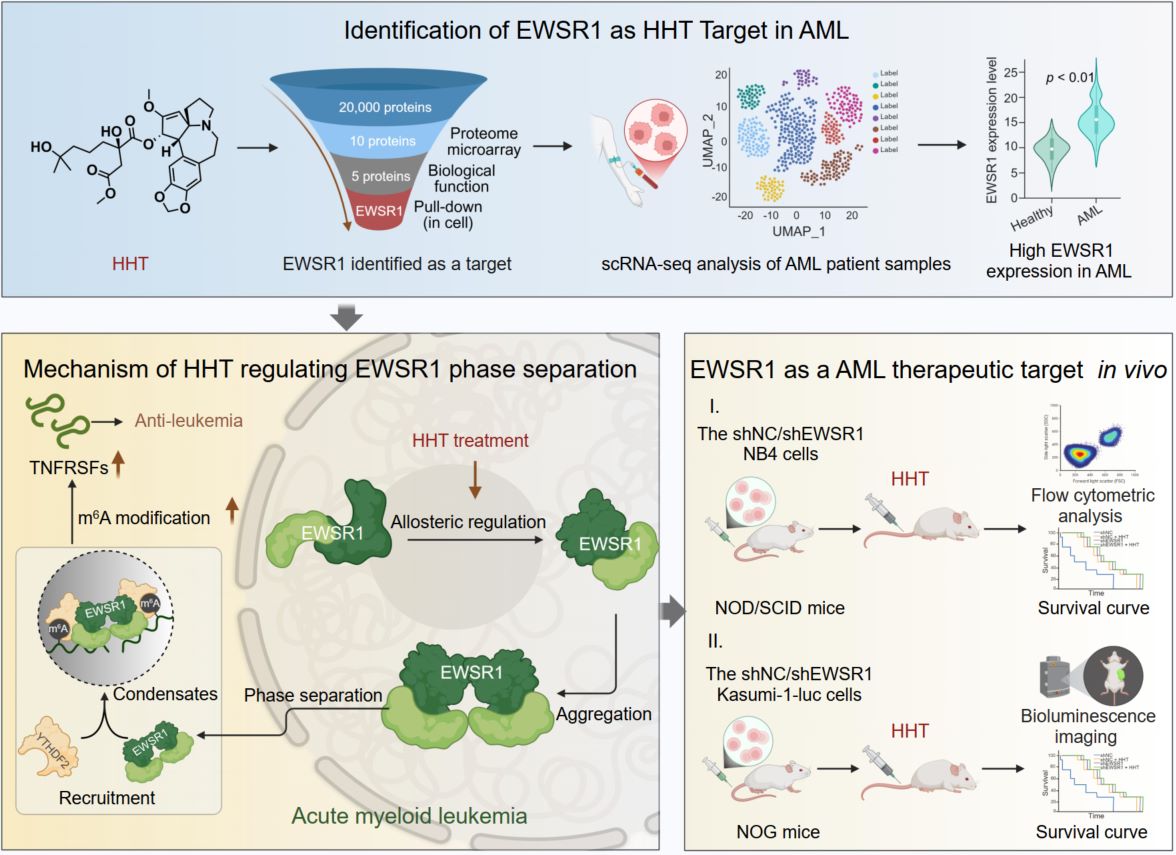
Homoharringtonine (HHT), a clinically approved anti-leukemic drug, directly binds the RNA-binding protein EWSR1. HHT induces an allosteric conformational switch in the RNA recognition motif of EWSR1, driving self-association and liquid–liquid phase separation (LLPS). The resulting condensates act as regulatory hubs that sequester the m6A reader YTHDF2, thereby preventing recognition and decay of m6A-modified transcripts. Stabilization of apoptosis-associated RNAs such as TNFRSF1B and HMOX1 disrupts leukemic cell homeostasis and suppresses AML proliferation. These findings identify EWSR1 as the direct effector of HHT and a predictive biomarker for therapeutic responsiveness in AML.
A comprehensive benchmarking for spatially resolved transcriptomics clustering methods across variable technologies, organs, and replicates
- 09 October 2025
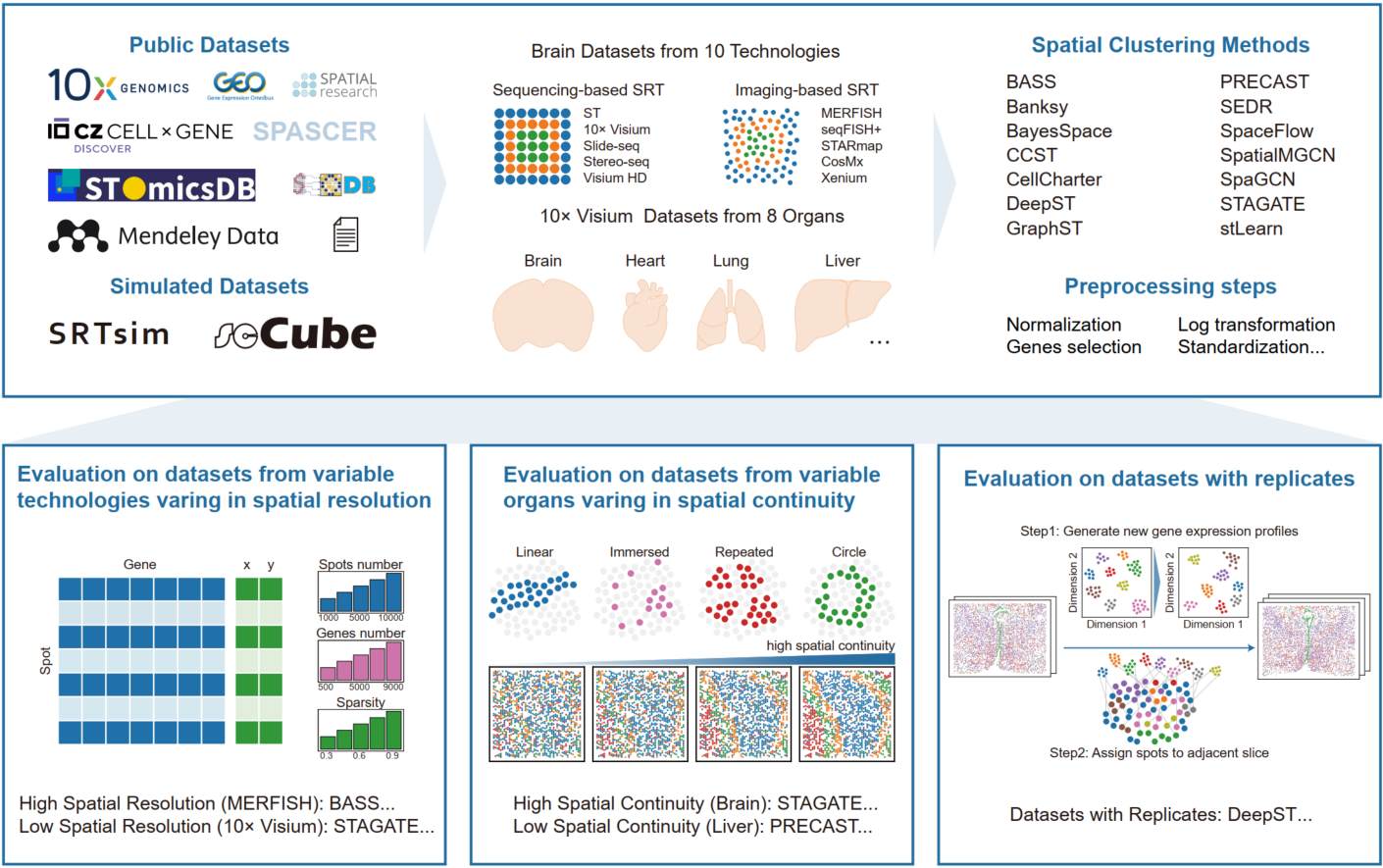
Spatial clustering is a critical step in spatially resolved transcriptomics. Here, we conducted a comprehensive benchmarking analysis of 14 spatial clustering methods using ~600 datasets, including both real-world and simulated data spanning ten technologies and eight organs. We provide practical recommendations for method selection across diverse technologies, organs, and biological replicates. Performance differences are largely explained by data characteristics, spatial patterns, and preprocessing strategies, and we establish an optimized preprocessing pipeline to enhance clustering robustness.
The interplay between tissue-resident microbiome and host proteins by integrated multi-omics during progression of colorectal adenoma to carcinoma
- 04 November 2025
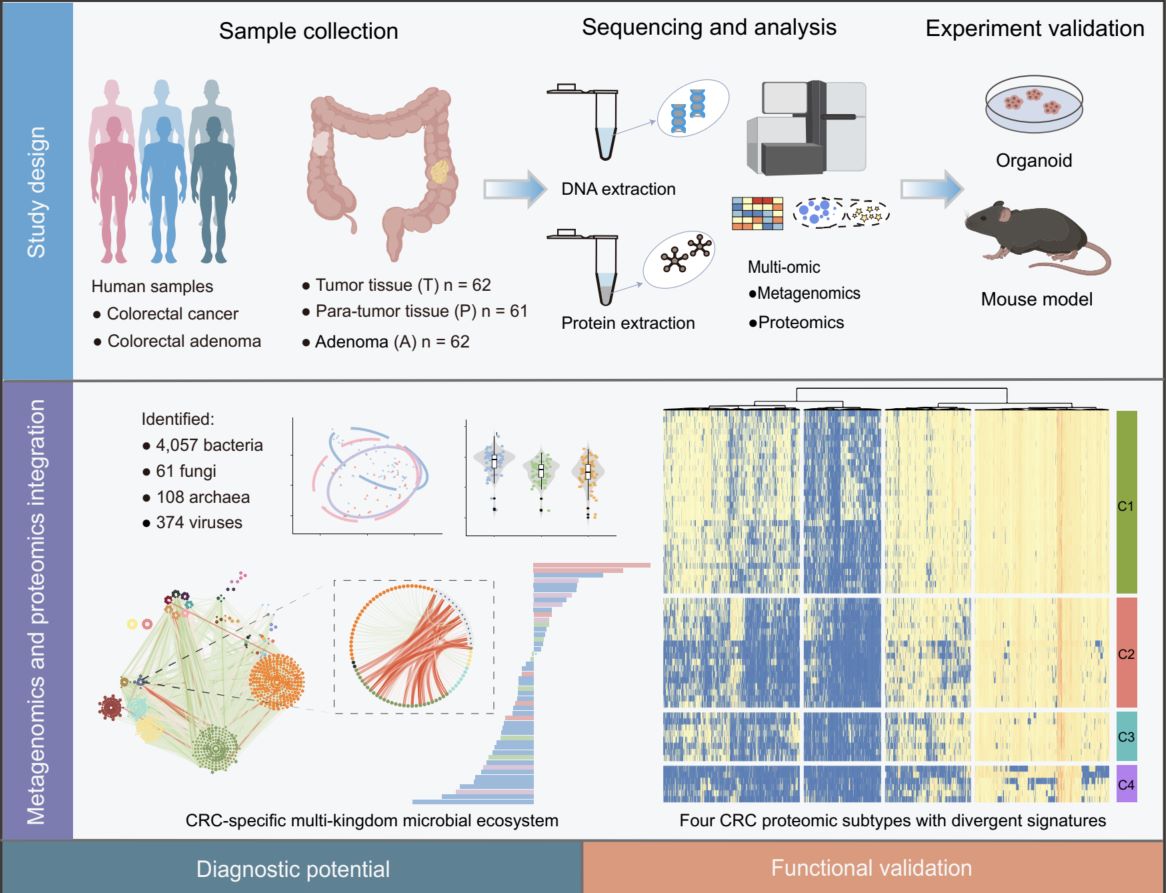
This study integrated metagenomic and proteomic profiling of 185 colorectal tissue samples—spanning adenoma (A), tumor (T), and para-tumor (P)—to characterize multi-kingdom microbiome and host protein dynamics in colorectal cancer (CRC). In total, 4057 bacterial, 61 fungal, 108 archaeal, and 374 viral species were identified, revealing CRC-specific microbial dysbiosis. Proteomic analysis defined four molecular subtypes (C1–C4) with distinct clinical outcomes. This study further developed a diagnostic model based on 14 microbial and 8 protein markers, which robustly distinguished adenoma from tumor (achieved an area under the receiver operating characteristic curve (AUROC) = 0.962) and early from advanced stages (AUROC = 0.926), with validation across multiple independent cohorts. Functional assays in patient-derived organoids and murine allografts confirmed that enterotoxigenic Bacteroides fragilis and Fusobacterium nucleatum promote tumor growth through Wnt/β-catenin and NF-κB pathway activation. These findings highlight dynamic host–microbiome interactions in CRC progression and provide novel biomarkers and therapeutic targets.
PhyloSuite v2: The development of an all-in-one, efficient and visualization-oriented suite for molecular dating analysis and other advanced features
- 25 November 2025
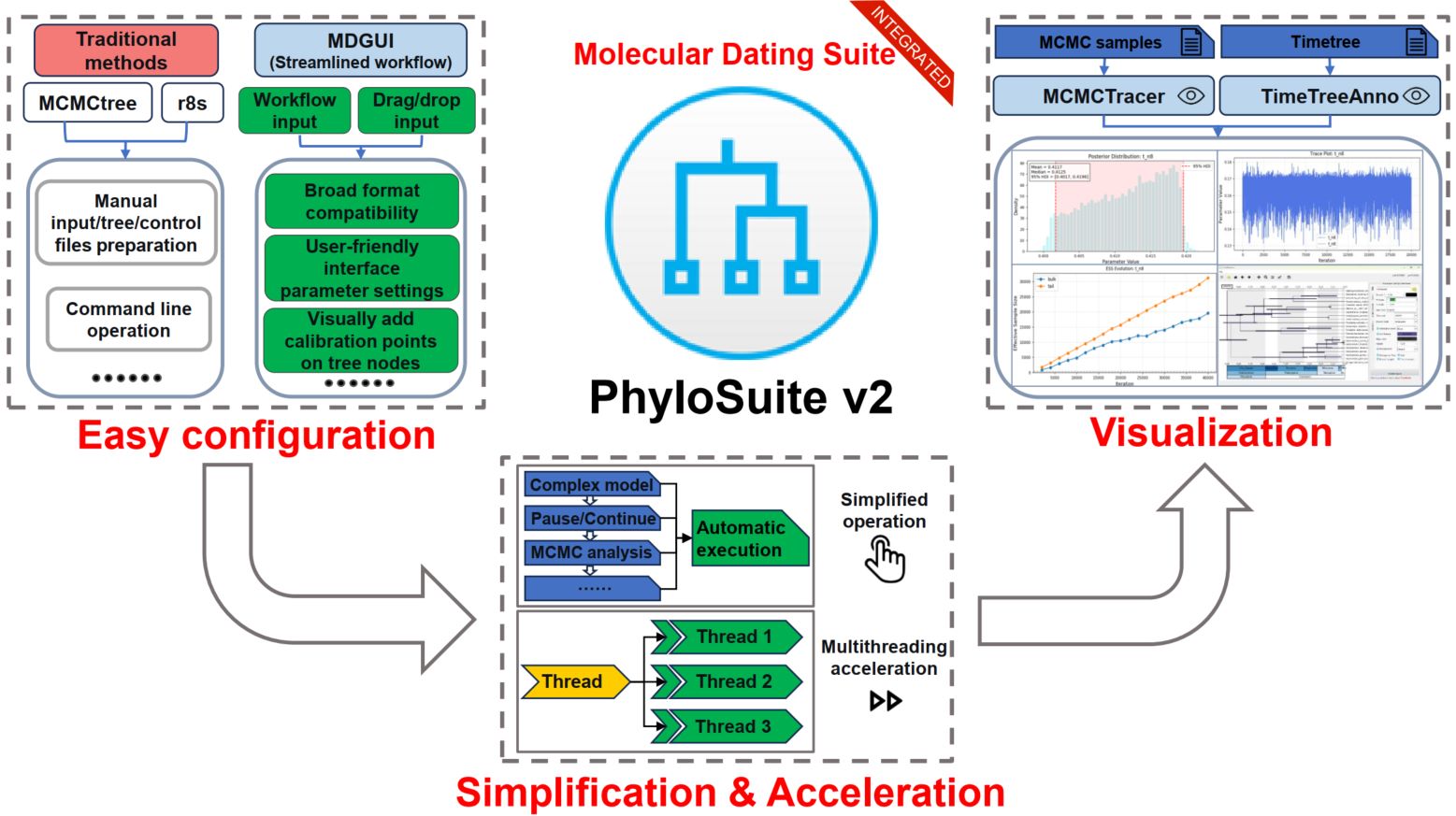
MCMCtree and r8s are among the most popular molecular dating tools in the current genomic era, but their utility is hampered by steep learning curves, particularly concerning input file formatting, the complexity of fossil calibration setup, tree visualization, and model selection. To enhance their usability and improve research efficiency, we developed three new tools: MDGUI (for molecular dating analysis), TimeTreeAnno (for timetree visualization), and MCMCTracer (for convergence assessment). We integrated these into the PhyloSuite v2 platform, along with MCMCtree and r8s plugins, to create a comprehensive molecular dating suite. Compared to existing solutions that we benchmarked, our toolkit offers a more intuitive interface and streamlined workflow, featuring visual calibration point configuration, support for multiple alignment formats, automated model selection and implementation for downstream analyses, one-click pause/resume functionality, multithreading acceleration, and on-demand MCMC convergence assessment and plotting. Furthermore, PhyloSuite v2 introduces other advanced features, including gene duplicate resolution during the extraction step, significantly accelerated data handling capabilities (specifically, format conversion and concatenation), deeper integration of the latest IQ-TREE models and functions, and further streamlining of the entire phylogenetic analysis workflow. The update also includes adaptation to high-resolution screens and numerous bug fixes. The source code for the new version of PhyloSuite is available at https://github.com/dongzhang0725/PhyloSuite.
Genome-wide CRISPR screen reveals an uncharacterized spliceosome regulator as new candidate immunotherapy target
- 28 November 2025
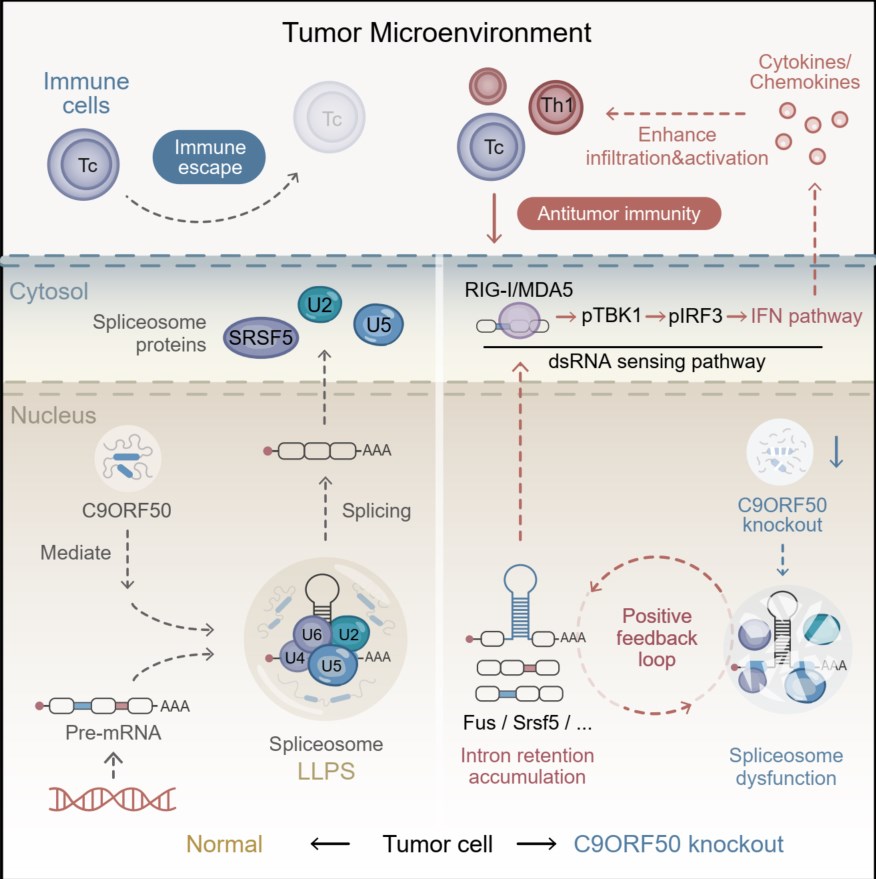
This study identifies the uncharacterized gene C9ORF50 as a novel regulator of immune evasion. Functioning as an intrinsically disordered protein, C9ORF50 drives liquid-liquid phase separation to facilitate spliceosome assembly and maintain RNA splicing fidelity. Genetic ablation or therapeutic inhibition of C9ORF50 disrupts this process, leading to widespread intron retention in pre-mRNAs of core spliceosome components. These unprocessed introns form immunogenic double-stranded RNA (dsRNA) that accumulates in the cytoplasm. This dsRNA is sensed by innate immune receptors (RIG-I/MDA5), activating the TBK1-IRF3 axis and subsequent type I interferon signaling. Consequently, this enhances the production of T-cell chemoattractants (CXCL9/10/11), promoting robust CD8⁺ and CD4⁺ T cell infiltration into tumors. Together, these findings reveal that C9ORF50 inhibition effectively converts immune-suppressive tumors into T cell-inflamed lesions, thereby enhancing immunogenicity and suppressing tumor growth.
Cultivar-specific preference of bacterial communities and host immune receptor kinase modulate the outcomes of rice–microbiota interactions
- 16 December 2025
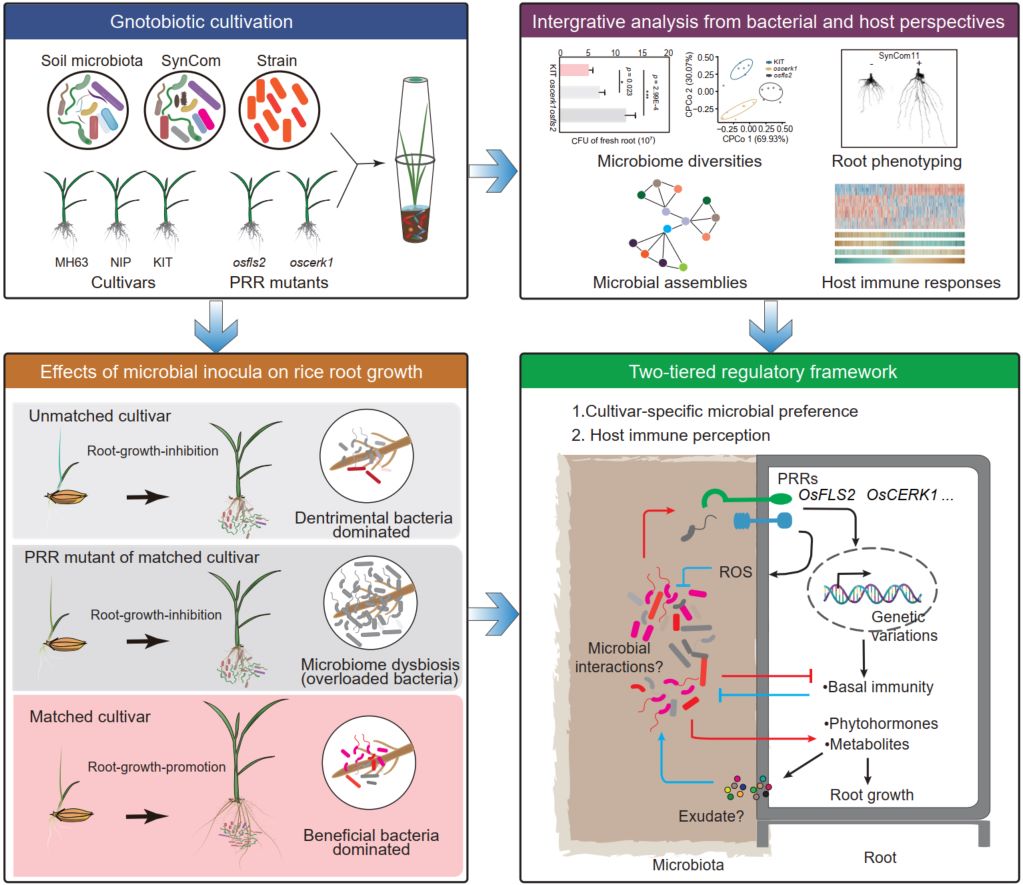
This study employed a tailored gnotobiotic cultivation system to investigate rice–microbiota interactions from both microbial and host perspectives. We found that the same bacterial inocula, including native soil microbiota, synthetic communities, and individual strains, exerted beneficial, neutral, or detrimental effects on rice root growth across cultivars. Our analysis suggests that a two-tiered regulatory framework, comprising cultivar-specific preference of the bacterial microbiota and host immune perception, governs the outcomes of rice–microbiota interactions.
Antioxidants promote metabolic remodeling in cattle rumen epithelium revealed by single-cell resolution
- 29 December 2026
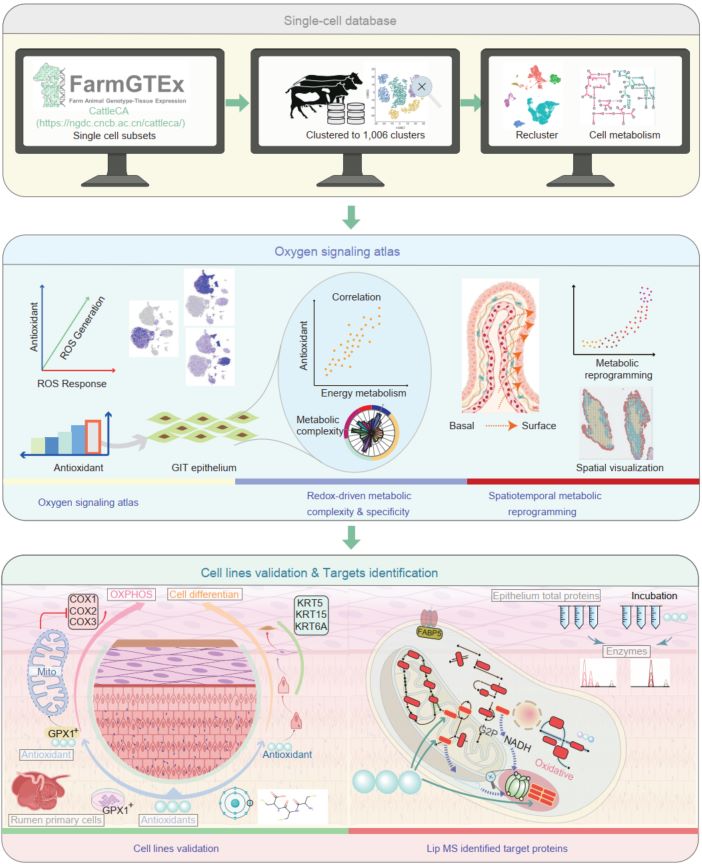
Single-cell and transcriptomics from 1.79 million cells identify the forestomach epithelium as the dominant site of antioxidant activity and oxidative phosphorylation (OXPHOS) in the bovine gastrointestinal tract. Along the basal-to-luminal differentiation axis, antioxidant capacity and OXPHOS rise in a coordinated metabolic–redox gradient. Primary rumen cell assays show that this gradient is strengthened by Cys-Cys-Cys supplementation or modulation of GPX1 expression, demonstrating a direct causal influence on epithelial metabolism. Key redox-metabolic regulators, including GPX1 and GSTP1, mediate these responses. Overall, antioxidants actively drive metabolic remodeling and promote epithelial differentiation in the cattle forestomach epithelium.
Metagenomics and digital cell modeling facilitate targeted high-throughput sorting of anaerobic hydrogen-producing microorganisms
- 21 September 2025
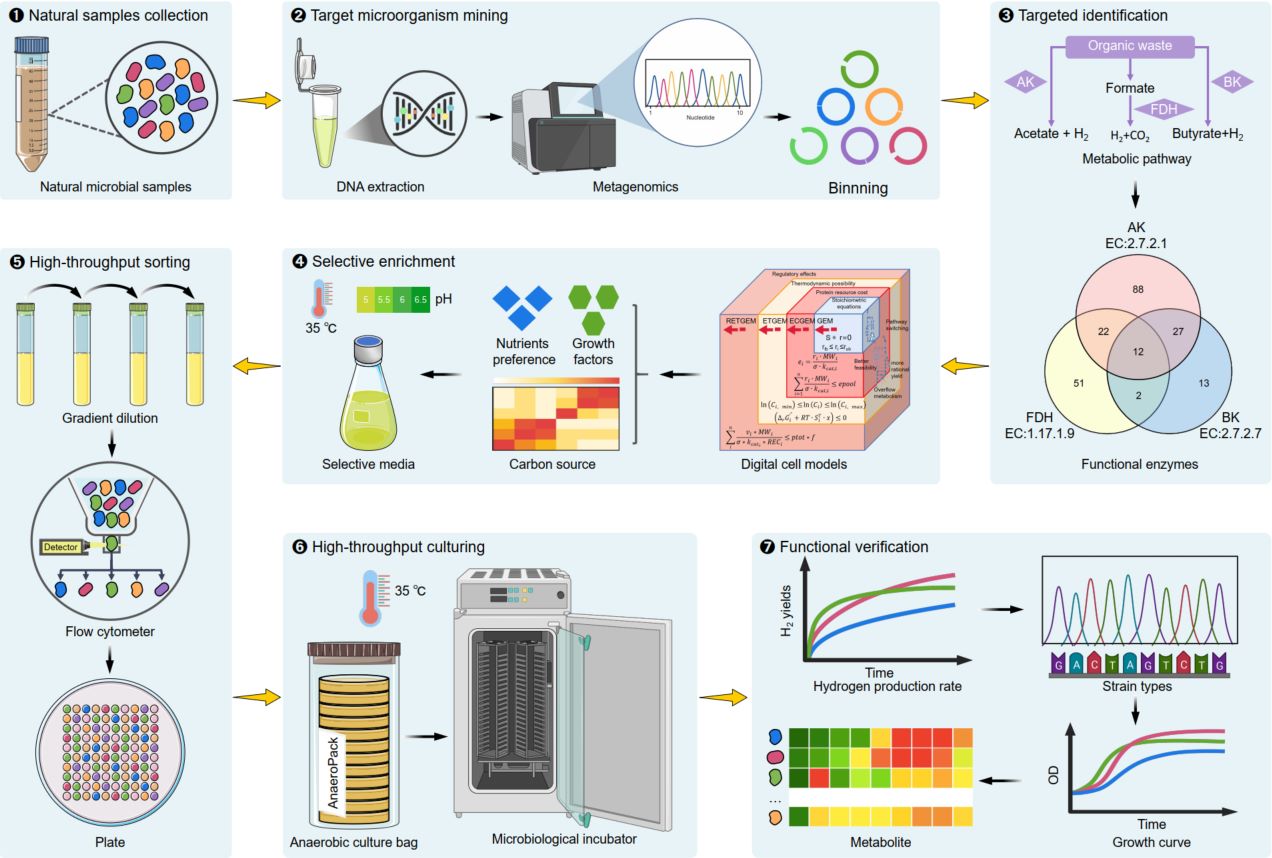
This study proposes a novel strategy that prioritizes functional recognition, followed by targeted high-throughput sorting, to enable the comprehensive, rapid, and efficient acquisition of target microorganisms. Using metagenomic sequencing and binning analysis, we identified 215 potential anaerobic hydrogen-producing strains from 12 large-scale biogas samples. Digital cell models were subsequently constructed from metagenome-assembled genomes, which guided the design of 14 selective culture media for enriching these hydrogen-producing bacteria. Flow cytometry-based high-throughput sorting successfully isolated 81 potential anaerobic hydrogen-producing strains, achieving a target acquisition rate above 37% and a survival rate exceeding 70%. This method holds broad potential for the discovery and sorting of functional microorganisms across diverse environments and may ultimately facilitate the development of synthetic microbiomes for industrial applications.
Nationwide profiling of vaginal microbiota in Chinese women reveals age-dependent shifts and predictive biomarkers for reproductive health
- 23 October 2025
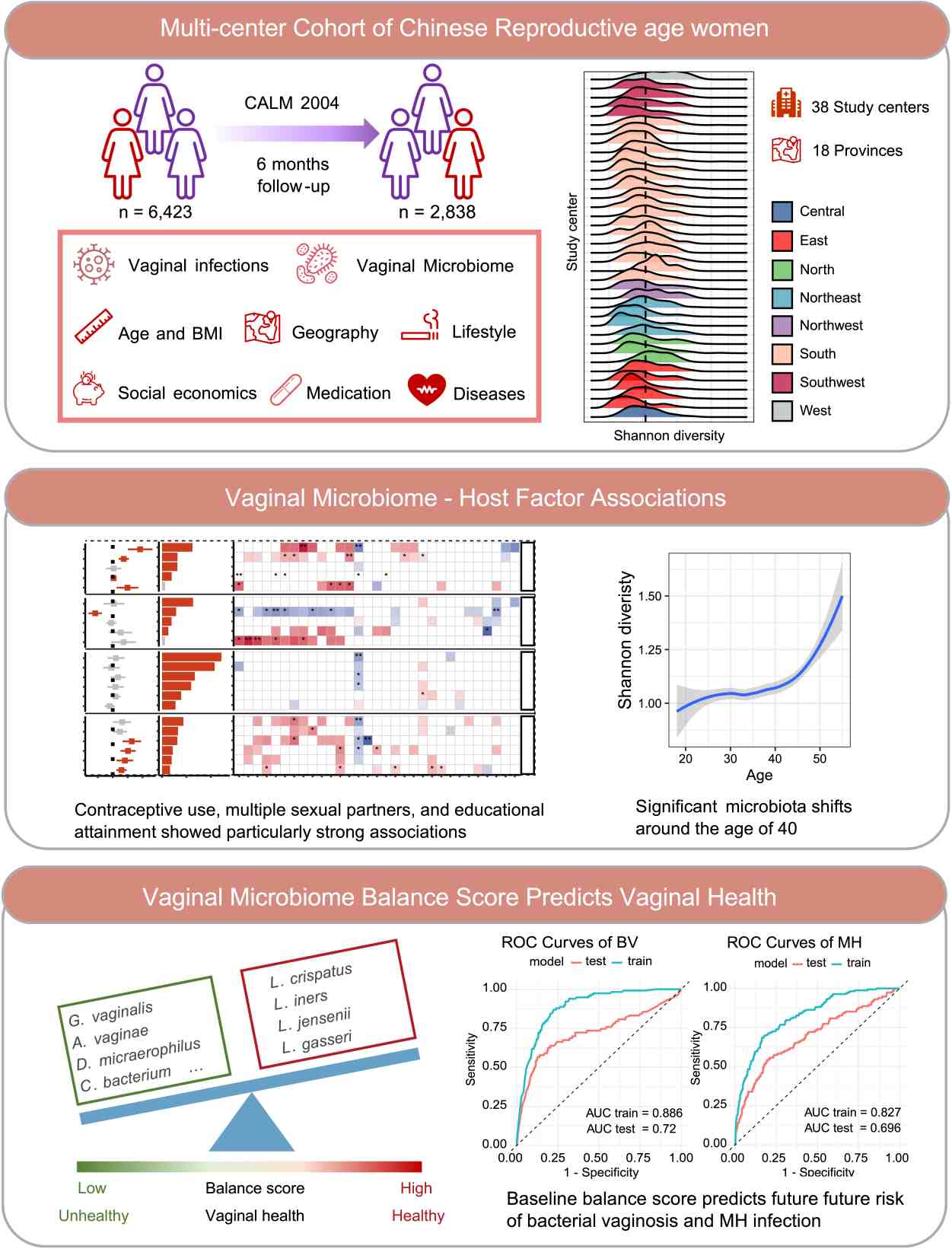
The vaginal microbiome is central to reproductive health, yet large-scale studies in East Asian populations remain scarce. Here, we characterized the vaginal microbiota of 6423 Chinese women of reproductive age across 18 provinces and assessed associations with 33 host factors. We observed a striking compositional transition around age 40, marked by declining Lactobacillus crispatus and enrichment of dysbiosis-associated taxa including Gardnerella vaginalis, independent of lifestyle or sociodemographic influences. Sexual behavior, contraceptive use, and educational attainment emerged as key determinants of community structure, differentially shaping Lactobacillus crispatus and Lactobacillus iners. Despite these associations, host factors explained less than 2% of overall variation, highlighting the resilience and individuality of the vaginal microbiome. To quantify vaginal health, we derived a microbiome balance score, validated it in external cohorts, and demonstrated its predictive power for incident bacterial vaginosis and sexually transmitted infections. Our findings establish a national-scale reference for the vaginal microbiome in Chinese women, reveal a midlife inflection point in microbial composition, and introduce a clinically actionable metric for risk stratification. These insights advance mechanistic understanding of host–microbiome interactions and inform strategies for precision interventions to preserve vaginal health.
Whole microbiota transplantation restores gut homeostasis throughout the gastrointestinal tract
- 11 November 2025
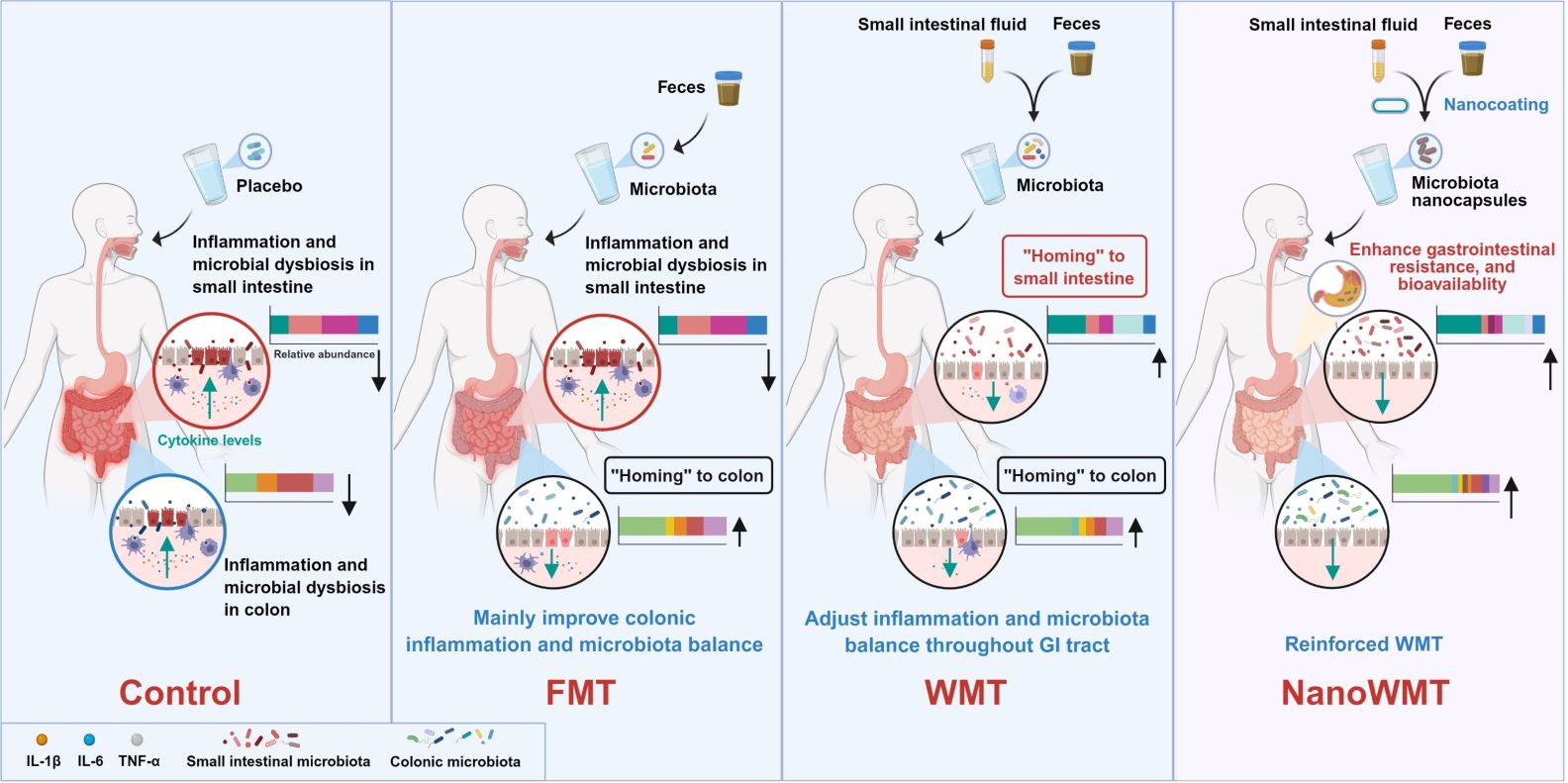
This study introduces whole microbiota transplantation (WMT), a synergistic therapeutic approach that concurrently transplants small intestinal and fecal microbiota. In germ-free mice, WMT outperforms conventional fecal microbiota transplantation (FMT) in restoring gut microbiota diversity and abundance. Moreover, in a chemotherapy-induced intestinal mucositis model, WMT alleviates intestinal inflammation and reverses microbiota dysbiosis. Encapsulation in layer-by-layer self-assembled nanocapsules further boosts microbial survival and colonization, amplifying WMT's anti-inflammatory effects and microbiota restoration in a mouse model of pan-intestinal infection. Overall, WMT represents a precise strategy for reshaping microbial homeostasis across the entire gastrointestinal tract, with therapeutic promise for inflammatory bowel diseases and small-intestinal disorders.
Soil-borne legacy facilitates the dissemination of antibiotic resistance genes in soil–plant continua
- 23 November 2025
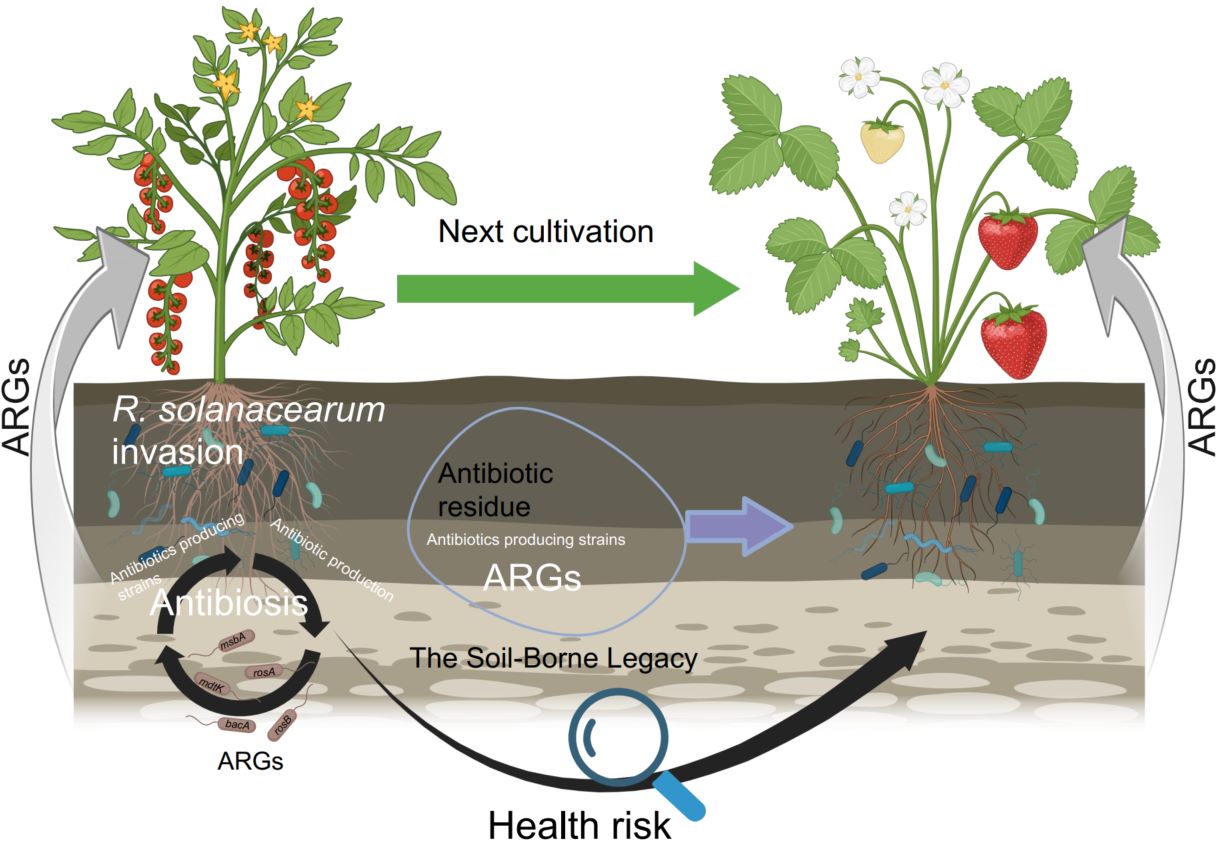
Antimicrobial resistance (AMR) disseminates throughout the soil–plant continuum via complex microbial interactions. Plants shape root- and leaf-associated microbiomes that sustain plant health; however, soil-borne legacies—enriched with antibiotic-producing microbes and resistance genes—govern AMR dynamics across agroecosystems. Using 16S rRNA gene sequencing, shotgun metagenomics, and high-throughput quantitative PCR, we profiled antibiotic resistance genes (ARGs), mobile genetic elements, and virulence factor genes across bulk soil, rhizosphere, phyllosphere, and root endosphere within soil–tomato and soil–strawberry continua. Recurrent bacterial wilt amplified the resistome, particularly polypeptide resistance genes, thereby establishing the rhizosphere as a major hotspot of ARG accumulation. Multidrug-resistant Ralstonia solanacearum (R. solanacearum) strains acted as major ARG reservoirs, harboring resistance determinants on both chromosomes and megaplasmids. Collectively, these findings demonstrate that pathogen-driven restructuring of the plant microbiome accelerates ARG dissemination, establishing soil-borne diseases as critical amplifiers of AMR across agricultural ecosystems.
A vast stem-progenitor cell pool, richly vascular system, and hybrid ossification drive the daily centimeter-scale elongation of bony antlers
- 08 December 2025
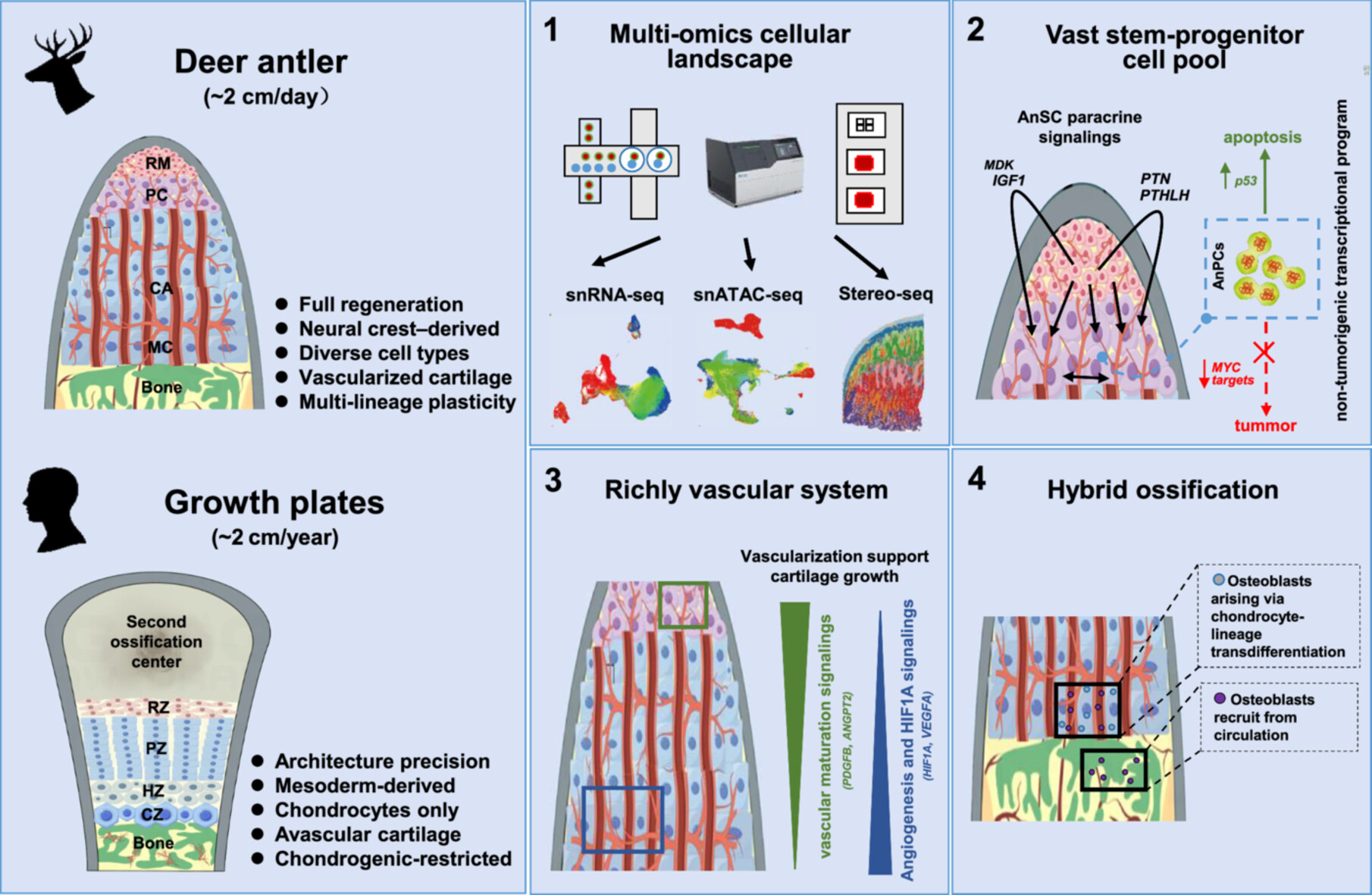
Bone growth and regeneration remain major clinical challenges. Deer antlers, the fastest-growing mammalian bone, regenerate via endochondral ossification and elongate up to 2 cm per day, far surpassing the ~2 cm annual growth of human growth plates. Here, we systematically mapped the cellular landscape of the antler growth center (AGC) using single-nucleus RNA sequencing, chromatin accessibility profiling, and spatial transcriptomics. The AGC harbors a large stem-progenitor pool that drives rapid elongation through vigorous proliferation supported by paracrine signaling. These proliferative cells exhibit a transcriptional program with intrinsically low tumorigenic potential, associated with apoptotic regulation. The AGC also establishes a vascularized niche that supports robust angiogenesis, sustains accelerated cartilage growth, and enables efficient recruitment of osteogenic cells. Notably, antlers employ a hybrid ossification strategy, combining endochondral ossification with direct hypertrophic chondrocyte-to-osteoblast transdifferentiation, likely via PHEX⁺ intermediates. Collectively, these findings refine fundamental concepts of endochondral ossification and offer insights for regenerative bone therapies.
Single-cell spatial transcriptomics reveals potential molecular mechanisms of Abelmoschus manihot (L.) medic in treating diabetic kidney disease
- 08 December 2025
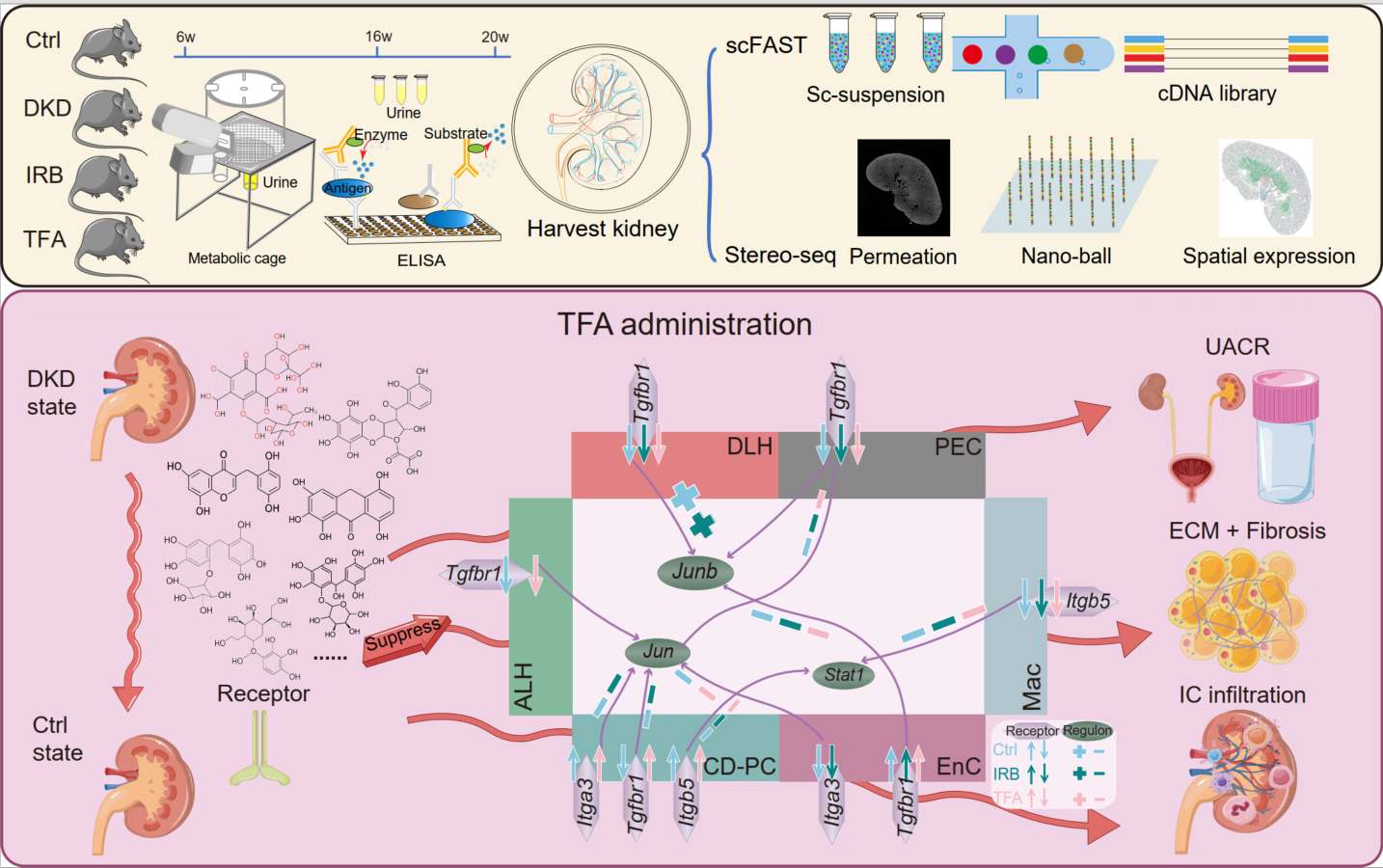
A clinical study reported that Abelmoschus manihot (L.) Medic (A. manihot), in the form of Huangkui capsule (HKC), combined with irbesartan (IRB) is an effective therapy for patients with diabetic kidney disease (DKD). The bioactive components of HKC are total flavones extracted from A. manihot (TFA). To explore the pharmaceutical molecular mechanisms underlying the efficacy of A. manihot in the treatment of DKD, we have combined SpaTial Enhanced REsolution Omics-sequencing (at 0.25 μm resolution) with single-cell full-length RNA sequencing. We employed the db/db mouse model of type 2 diabetes and DKD. These experimental methods generated the first single-cell resolution pharmacopathological spatial atlas in kidneys of db/db mice that were treated with TFA or IRB. TFA exhibited therapeutic effects on DKD comparable to those of TFA combined with IRB. Following genome-wide gene screening and molecular docking simulation, we have identified the key renal receptors (Itga3, Itga5, Tgfbr1, etc.) and regulators (Jun, Junb, Stat1, etc.) underlying the therapeutic action of TFA in DKD. This study provides novel insights into the pharmaceutical mechanisms of A. manihot in the treatment of DKD.